The Hot Spring Hypothesis for the Origin of Life and the Extended Evolutionary Synthesis
by Bruce Damer
8 May 2019

Origin of life researcher, Professor Bruce Damer, presents an exciting four-part essay on the Hot Spring Hypothesis for the Origin of Life, and how this connects to the Extended Evolutionary Synthesis.
Introduction to the Series
Part 1. The Origin of Evolution
Part 2. Programming Without a Programmer
Part 3. The Quest to Test (or Falsify) the Hot Spring Hypothesis
Part 4. The Progenote, Chickens, Eggs and the EES
Summary
Introduction to the Series
For the first time since Charles Darwin mused back in 1871 that life might have begun in a warm little pond, scientists have proposed and are testing an experimentally accessible hypothesis for life’s beginnings… in a hot little cycling pool which would have existed on a volcanic land mass on the early Earth. This series of articles will review the first decade of work on the hot spring hypothesis for the origin of life and cover its early empirical testing in the laboratory, at field analog sites and speculate on its broader implications including the search for life beyond the Earth. Most pertinent for readers following the development of the Extended Evolutionary Synthesis (EES) is that one of the core insights that the hypothesis implies is that life’s start and early evolution was made possible by a form of proto-niche construction – that is, the construction of life-promoting environmental conditions by non-living pre-cellular entities – in concert with a collaborative network effect. This new idea is in direct opposition to the more commonly held notion that all of life (including its origins) must arise through competition between individuals and the passing of genetic traits solely down through individual species lineages.
Perhaps the greatest construct of the human intellect is our surprisingly predictive mathematical formulations describing cosmogenesis, the origin of the universe and the operation of its physical laws. Twenty-first Century science has now turned its focus to another unresolved origin question, biogenesis, seeking approaches to the mysterious process which transmuted inanimate matter into the living world. Life as we know it operates universally through the unit of the cell, a membrane-bounded compartment which crowds together systems of interacting polymers. These polymers and their compartments are fashioned by highly complex molecular machines. The conundrum which faces origin of life investigators is how such sets of polymers could first arise without these machines to make them. This is the primordial version of the age-old question: which came first, the chicken (polymers) or the egg (the molecular machines which make them)? In addition, because all living entities are cell based, and the Earth’s earliest fossil record revels communities composed of single celled organisms, we may have been misled into assuming that life originated as self-sufficient single cells. Self-sufficient cells require a high degree of evolutionary technology so we are forced to consider a new scenario: that life originated as networks of much simpler protocells that collectively constructed a proto-niche able to support growth and evolution.
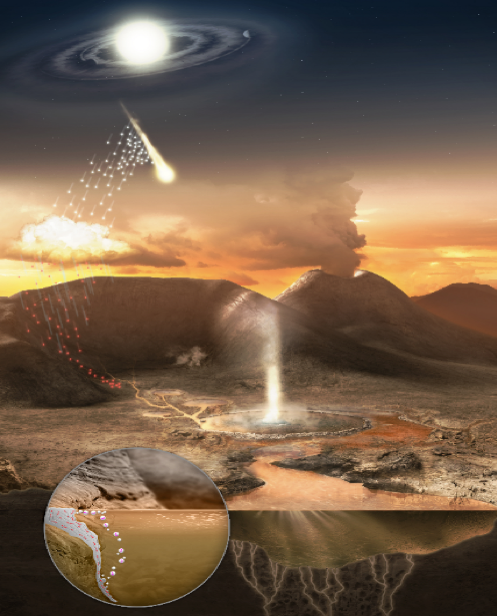

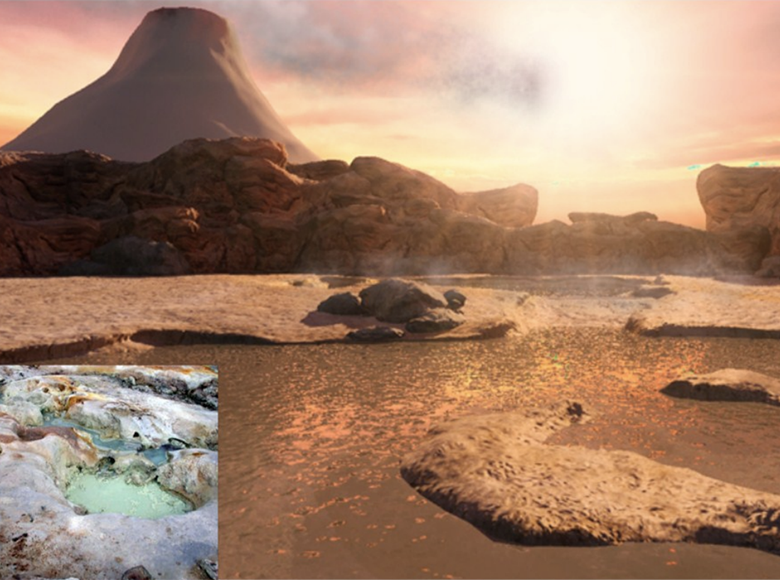
Figure 1. View of the Hadean Earth four billion years ago. Image: Ryan Norkus and Bruce Damer.
Join us now for a journey back four billion years ago to a very different Earth (figure 1), where the skies were orange-brown with carbon dioxide and volcanic gases and lit by frequent flashes of incoming meteorites from the dusty disk of the still-forming solar system. A great global ocean, brown with dissolved iron and other minerals, roiled with tsunamis and gigantic tides pulled by a moon five times closer than it is today. Rising from the oceans were numerous roaring volcanoes that were just beginning the billion-year process of radiating the Earth’s formation mantle heat through its new, thin crust. Lava flows poured down the flanks of the volcanos, building up great land masses onto which storms dumped deluges of fresh rainwater. This water found its way underground, infusing chambers which boiled the water and erupted it back up to the surface, driving immense bubbling hydrothermal fields. In one field a steaming geyser rose up like the greatest granddaddy of Old Faithful, flooding its hot, enriched waters out onto the basaltic plain. Thousands of pools filled and chemically interacted with these infusions of hot water. Tiny grains of dust as well as large meteorites which had formed in the great flattened accretion disk of the Solar System were vacuumed up by the Earth and fell on this landscape. As the dust and meteorite fragments blew or were washed across the landscape, they collected in freshwater pools, some of them connected to active hydrothermal springs and dissolved their loads of precious organic compounds into the pools. As the pools dried down, they became surrounded by a glistening bathtub ring composed of layered membranous structures formed by fatty acids delivered by the meteoritic material. Between those silvery drying layers, the organic building blocks of amino acids and nucleic acids jostled into position. The first primitive biopolymers were forming, and our origin of life story is now beginning.
Part 1. The Origin of Evolution
Charles Darwin’s Theory of Evolution can be expressed succinctly in these four points: 1) individuals of a species have largely similar but not identical traits; 2) their traits are passed by reproduction to subsequent generations; 3) more organisms are reproduced than can survive; and 4) only the survivors from the resulting competition for resources will go on to reproduce. Variations of the traits of individuals give some members of the species advantages in cycles of competition and reproduction, passing these traits on to the next generation. A species could be defined as a population of organisms sharing a nearly identical set of traits. An organism which is a member of a species is by definition a fairly complex thing, with a distinct genotype and the behaviors and products expressed by them, the phenotype. Even simple cellular organisms such as a bacterium, are witheringly complex molecular machines. Yet at life’s origins there were no complex systems to support distinct species and in fact, no self-sustaining protocells; only relatively simple structures and interactions would have been possible. How simple? Just good enough to permit a system to grow and evolve. To evolve there must have operated some natural, already available process that could drive the earliest form of proto-evolution. After a long process fraught with frequent failures and restarts of uncountably large numbers of simpler entities, the first living cells capable of dividing and passing on accumulated traits would emerge. The unit of evolution that gave rise to these living cells must have initially been entirely passive, completely dependent on self-assembly in an environment of physically and energetically driven cycles fed by externally supplied building blocks. This first part of the series will propose how an initial ancestral network of interacting molecules kick-started the evolutionary process and gave rise to the living world.
The geochemical underpinnings of evolution
Underlying Darwin’s definition for evolution by natural selection, the simple act of staying alive long enough to reproduce is fundamental. That requirement can be rephrased as the sustained maintenance of an away-from-thermodynamic-equilibrium system. For all of life on Earth, membranes encapsulate systems of interacting polymers which are constantly undergoing repair and re-synthesis. The system utilizes sources of energy to maintain itself in an equilibrium which is far from a state of a random soup of similar chemicals. Through generations, the system adds complexity: a new gene here, a new structural or behavioral expression there. This can also be referred to as a chemical ratchet or kinetic trap, in which the rates of synthesis exceed the rates of degradation (Hargrave et al. 2018; Higgs, 2016; Ross and Deamer 2016). Death and subsequent decay could simply be defined as a system of molecules traveling in the reverse direction in a kinetic trap.
Equipped with this key underpinning, we can now tackle the question: how did evolution itself originate? For the answer we shall take an informed conceptual journey back into deep time. We must find a plausible natural setting and mechanism that would have given rise to the progenote, the product and subject of these first steps of evolution.
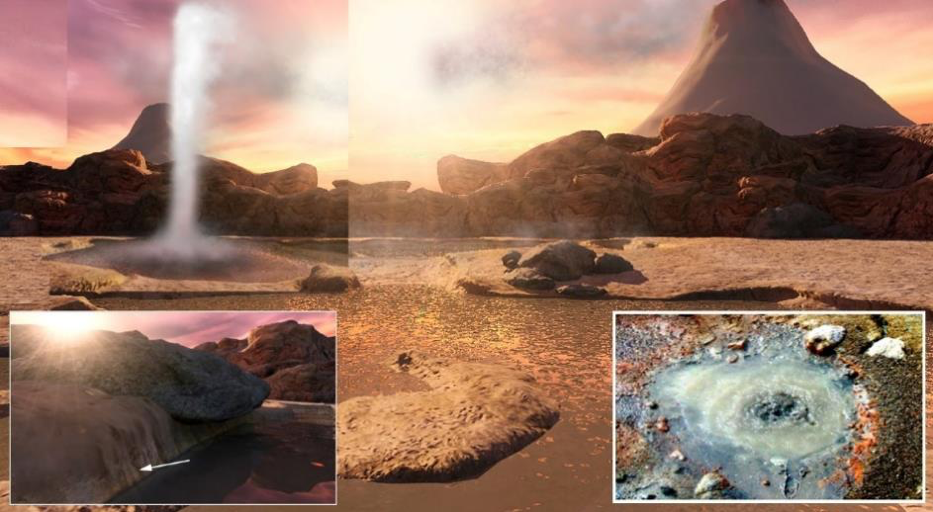

Figure 2. An artist’s conception of a Hadean volcanic landscape in which cycles of evaporation and rehydration can occur, driven by periodic geyser action. Inset (lower left) shows a ring of dried solutes on the mineral surfaces at the edge of a fluctuating pool. Inset (lower right) shows a boiling pool associated with a hot spring site on Mount Mutnovsky in Kamchatka, Russia. Image: Bruce Damer; photo: Tony Hoffman.
What chemical, geological and temporal setting would reliably sustain a purely chemical (pre-biological) kinetic trap? The environment must be bounded to contain and concentrate chemicals and cycled regularly with a continuous source of energy and steady supply of useful building blocks. On the early Earth at the time estimated for life’s beginnings approximately four billion years ago, there was only one such environment with these properties: hot spring pools associated with volcanic activity. Figure 2 depicts such a pool driven by regular refilling by a nearby geyser. Distant volcanos build new landmasses which collect materials sourced from above – including material falling in from space, as well as precipitates from volcanic gases and atmospheric chemistry – and also delivered from below through hot spring sources.
The presence of organics on asteroids and comets as demonstrated by recent space missions has created a solid case that the dusty accretion disk of the early solar system delivered meteorites and tiny dust particles rich in organic compounds including fatty acids, amino acids, sugars, and nucleobases to the early Earth (Pearce et al, 2017). If these fell into the oceans they would be lost in the dilute bulk, unable to participate in the chemistry for life’s beginnings. If they fell onto land they can be washed into pools, concentrating sufficiently to permit prebiotically essential chemistry to start.
Organics separating out from rocky meteoritic materials have been shown to form membranes from carboxylic (fatty) acids also known as lipids (Hargreaves and Deamer, 1978). Amino acids also enter solution along with nucleobases, building blocks which can form source monomers that can then link together into polymers. The formation of polymers in sufficient length to support catalytic activity is a prerequisite to the emergence of any living system.
Solving the ironic (and problematic) Water Problem of the origin of life
Perhaps the ultimate irony of the chemistry of life is that it takes place in water, an environment which is also very good at degrading polymers in a process called hydrolysis. Banged around by the motions of molecules of water, polymers can come apart. Life has managed to deal with this seeming contradiction with the evolution of complex enzymes that constantly synthesize, repair and then re-synthesize polymers in the presence of water. At life’s beginning phases there were no enzymes. So, for the polymers to first form, they must take advantage of the one mechanism that will allow them to form: dehydration synthesis. Polymerization of amino acids into peptides and nucleic acids into polymers like RNA and DNA are called condensation reactions as water is a leaving group. Decades ago it was established that polymers could form without enzymes in drying down mixtures subject to sufficient heat (Sawai et al. 1975). More recently our group under the direction of my colleague Prof. David Deamer demonstrated using experimental laboratory chemical simulations (figure 3) that not only could polymers form long chains in dehydrating conditions from two building blocks of RNA (AMP and UMP) but that these polymers could attain great lengths exceeding 100 units in the presence of lipid membranes that are known to form from fatty acids delivered by meteorites (Rajamani et al. 2008, Deamer et al. 2019).
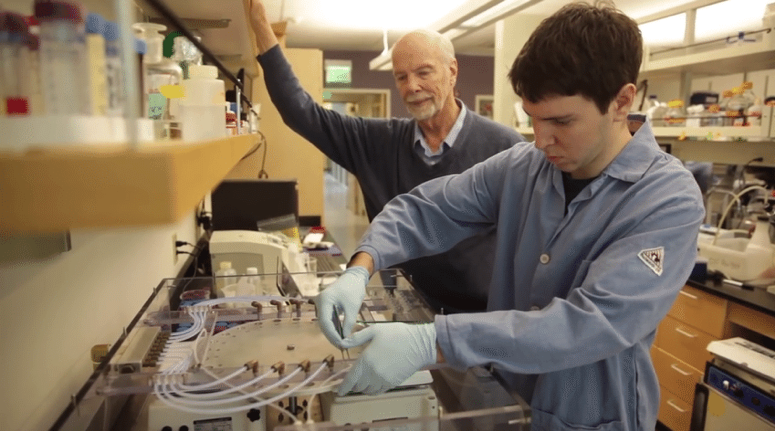

Figure 3. Professor David Deamer (left) with student Ryan Lorig-Roach (right) in the laboratory at UC Santa Cruz working with a wet-dry simulation chamber.
The most accessible dry place in a natural chemical laboratory available for life’s origination would be the mineral surface at the edges of pools, a place where today a “bathtub ring” of drying solutes is often observed (figure 2 insets). This drying area, which might also include pool bottoms if they dehydrate completely, provides polymers and other compounds a temporary refuge from the relentless degrading effects of hydrolysis. When these pools refill the bathtub rings will deliver their concentrated compounds, including polymers formed there, back into the bulk fluid of the pool.
Figure 4 illustrates how this occurs. First (bottom) a small, hot spring pool containing the building blocks and a membrane-forming compound such as a fatty acid or a mixture of different lipids dries down. The monomers are squeezed into a two-dimensional space between layers of membrane (inset: freeze fracture image of dried lipid lamellae). As water departs through the stack of layered membrane in the final stages of dehydration, the monomers jostle together and link, forming bonds. In figure 4 (top) when a pulse from the hot spring or another source of hydration fills the pool the outer membranous layers bud off large numbers of vesicles, some containing one or more randomly synthesized polymers (inset: stained microscope image of budding off vesicles, some containing DNA). We refer to vesicles containing polymers as protocells. The intermediate moist phase (middle) is a hydrogel of 50% water and lipid compartments (inset: microscope image of vesicles lining up in drying solution) formed as the pool dries down. Our protocells jostle together forming into a gel aggregate together with increasingly concentrated solutes. The protocells will then begin to fuse together into layers, returning their contents to another step of dehydration synthesis. Each of these three phases subjects populations of polymers to different selection factors: in the dry phase there is competition for monomers by polymers, in the wet phase selection acts for the stabilizing effect of polymers on the surrounding membranes that protect them from loss and degradation and, in the gel phase there is the opportunity to select for metabolic activity that might amplify populations of polymers in future cycles. This is the “coupled phases” scenario first described in (Damer and Deamer, 2015) and fully described in (Damer and Deamer, 2019) in which populations of polymers are coupled between these three phases, subject to combinatorial selection (before Darwinian natural section) and can undergo molecular evolution. Laboratory and field testing around the world have empirically demonstrated the first steps in this scenario (Deamer et al., 2019).
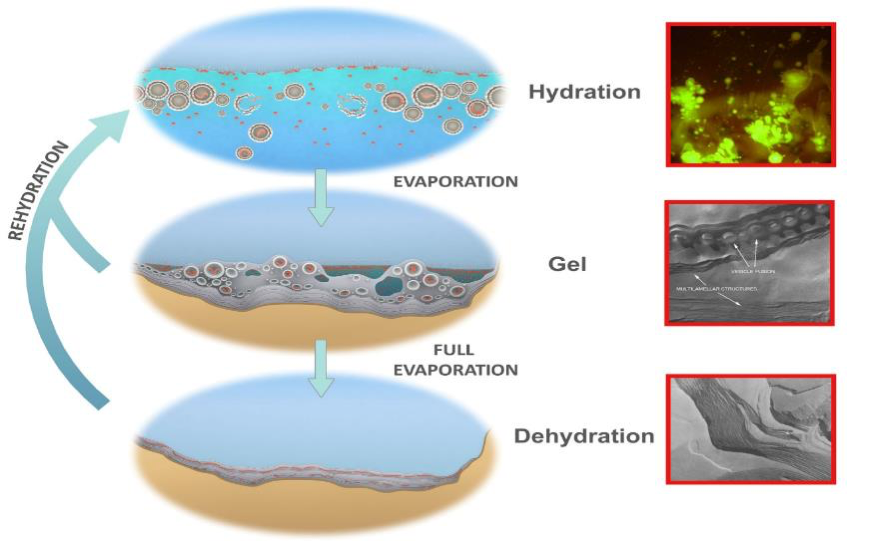
Figure 4. Natural drying and re-wetting cycles in a small pool supporting three phases of membranous encapsulation. Hydrated phase (top), inset: protocells containing DNA budding out of a dried mixture of DNA and phospholipid, stained with acridine orange. A moist gel phase (middle), inset: hydrogel of lipid with approximately 50% water by weight showing lipid vesicles fusing into lamellae. Dehydrated phase (bottom), inset: freeze fracture image of anhydrous lipid lamellae of phosphatidylcholine. Micrographs: David Deamer; Image: adapted from (Damer, 2016).
The Progenote and Proto-Niche Construction
The progenote, a transitional form which occupies the time frame during the origin of life between inanimate matter and living cellular systems, was first defined by (Woese and Fox, 1977; Woese 1998, 2002). While the actual form of the progenote is not specified, it is abstractly described as an entity engaged in the collective process of evolving the relationship between phenotype and genotype and is described more specifically by (Arnoldt et al., 2015):
Progenotes were not well-defined organisms as such, because they had no individuality and no long-term genetic pedigree. Their genes and component parts could come and go, being swapped in or out with other members of the community via horizontal transfer. But because biochemical innovations produced by any member of the community were available to all, evolution at this time was rapid—probably more rapid than at any time since. Selection acted on whole communities, not on individual progenotes. Those communities that were better at sharing their biochemical breakthroughs flourished. Out of this cauldron of evolutionary innovation, the universal genetic code and its translational machinery co-evolved in response to the selective pressures favoring efficient sharing and interoperability
The progenote we are proposing as a key component of this origin scenario begins as a passively assembled aggregation of “member” protocells which when subjected to a large number of selective cycles, evolve into the active, self-managed state of cellular life. We hope to show throughout these essays, that as these member protocells are subject to evolution through selection, they can begin to collectively forge and modify their environment, the temporary aggregate we term the progenote. Over time the progenote itself becomes subject to distribution, evolution, growth and adaptation. In Part 4 we will illustrate that some of this collective evolution will involve the emergence of a regulated network of chemical reactions as the products of metabolic reactions diffuse throughout the protocell aggregate. This regulation will also apply to protocell and progenote membranes. In this way we hope to make the case that such a progenote is in fact a plausible example of the first “proto-niche construction” and that niches are in fact central to life’s origins.
Putting it all together into the Hot Spring Hypothesis for an origin of life
In (Damer, 2016) and (Damer and Deamer, 2019) the scenario for a hot spring hypothesis for life’s origins was set into an early Earth landscape (figure 5). This diagram illustrates the entire scenario from the interplanetary synthesis and delivery of meteoritic inputs to a volcanic landscape (step 1), the accumulation and concentration of organic compounds to support the initial chemistry to create membranous vesicles (steps 2 and 3), the influence of a hydrothermally pulsed pool driving the three cycling coupled phases system to generate populations of protocells and form progenotes (step 4). Next come the pathways progenotes must travel, scaling evolutionary gradients to early living microbial populations through distribution and adaptation across a landscape (steps 5-6). Early life then engages in global colonization of both aqueous fresh water settings on the land and the more extreme intertidal and marine shorelines around landmasses (step 7). After a great number of additional evolutionary steps, highly robust microbial communities emerge which are advanced enough to leave fossil traces preserved as stromatolites. Recent stromatolite discoveries provide evidence of life thriving in hot springs in pools on land as far back as 3.5 billion years (Van Kranendonk et al. 2017; Djokic et al., 2017) as well as a solid fossil record of life in lake and marine shoreline environments.
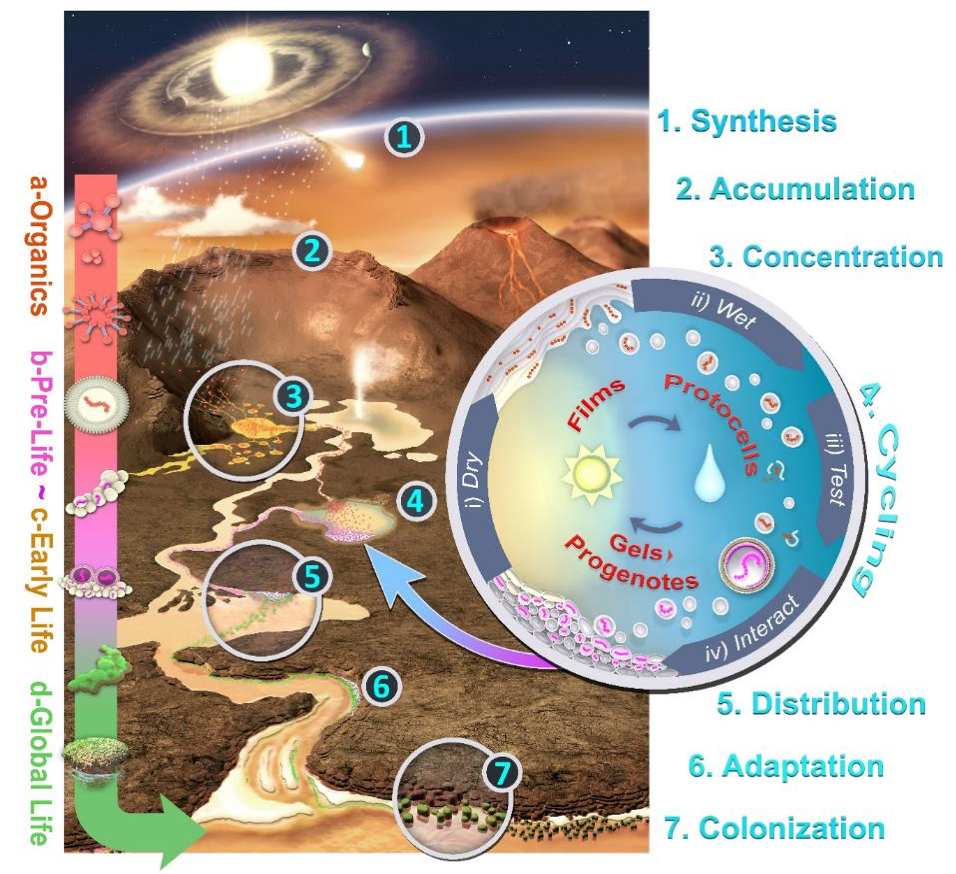
Figure 5. Integrating a prebiotic Hadean geological landscape with the chemistry of life’s origins in hydrothermal fields and its subsequent adaptive pathway from progenotes into early microbial communities. Image: Bruce Damer and Ryan Norkus (adapted from (Damer, 2016)).
To discover the origin of evolution we have first sought and identified a physical-chemical process, geological setting and unit of selection able to undergo a primitive form evolution, growth, distribution across a landscape, and adaptation through the scaling of selection gradients. The resulting hot spring hypothesis for an origin of life provides an experimentally testable instantiation of this process as well as the unit of selection: a progenote self-assembled from aggregates of protocells. These protocells are lipid encapsulated systems of interacting polymers cycling through three coupled phases in a periodically filling and drying hot spring pool. The first few steps of this hypothesis have passed empirical testing in the laboratory and in volcanic hydrothermal field analogs of the early Earth (Deamer et al. 2019). We will next address the proposal that at its origin and throughout its history, life on Earth has been primarily a project of niche construction suffused by a collaborative, networked sharing of resources driven by cycling energy sources. The next three essays will elaborate on and attempt to justify this proposal and address what can we apply from this model of the origin of life to a clearer understanding of its subsequent evolution throughout Earth’s history.
Part 2. Programming Without a Programmer
In Part 1 we took up the question of the origin of evolution, suggesting the underpinnings of the process described so succinctly by Charles Darwin in his book Origins of Species (1859). We identified the need for a kinetic trap operating on a self-assembled unit of selection: a population of protocells which undergo three phase changes, one of which temporarily forms a gel aggregate, the progenote. We then introduced a plausible pathway to the arising of the progenote through cycles of selection in hot spring pools on the early Earth. We put it all together by depicting a journey across a volcanic landscape which large populations of protocells and their progenote collectives might have taken to cross the evolutionary chasm to the first dividing, cellular life. We concluded by suggesting that life on Earth, from its origins and throughout its history, was primarily a process of niche construction paired with emergent collaborative networks.
In part 2 we will take up the task of convincing you that it is possible for the processes and structures of biology to arise de novo. In other words, that the scenario we are proposing can write the programs of the chemical operating system of life, without a programmer.
A new view into Darwin’s Warm Little Pond

Figure 6. Charles Darwin and his much-cited 1871 letter to J.D. Hooker.
In Part 1 we described how hot spring pools possess the chemical and thermodynamic properties which support the chemistry that can lead to life. Perhaps this should not surprise us as it was the father of evolutionary biology himself, Charles Darwin, who penned a remarkably similar and prescient vision in a letter to his friend J.D. Hooker (Darwin, 1871):
But if (& oh what a big if) we could conceive in some warm little pond with all sorts of ammonia & phosphoric salts,—light, heat, electricity &c present, that a protein compound was chemically formed, ready to undergo still more complex changes…
If we take his sentence apart with modern understanding, we agree that a little pond would provide molecules the chance to be concentrated enough for reactions to occur and warm enough water provides activation energy for those reactions. His light and electricity suggest sources of higher quality energy than simple heat to more precisely drive reactions. His ammonia and phosphoric salts would provide some of the building blocks to link together into a molecule known to Victorian science: proteins. His final statement that a protein compound was chemically formed, ready to undergo still more complex changes is where our part of the story begins. These words revealed that Darwin understood that proteins (polymer chains composed of monomer building blocks) must somehow be synthesized but then re-formed in still more complex configurations. This is what chemists today call an “away-from-equilibrium system” and is characteristic of all of life.
For the past century and a half after Darwin penned this famous phrase, investigators in the origin of life pursued several illuminating but ultimately dead-end chemical scenarios. These scenarios have been situated at many venues including: spark discharge chambers simulating an early Earth atmosphere (Miller, 1953), coacervate droplets investigated by Alexander Oparin and Sidney Fox, clay surfaces as an organizing matrix proposed by Cairns-Smith, and the possibility of prebiotically important reactions occurring at energy gradients in deep ocean hydrothermal vents (Russell et al. 2014). However, with little experimental evidence showing a prebiotically-plausible pathway to Darwin’s ever-complexifying protein compounds, workers in the field have increasingly returned their focus to the venue described in his letter: the warm little pond.
Until the past decade, a way forward was not visible to most chemists because they are used to working with solutions, that is, wet test tubes of compounds in which reactions occur. The “water problem” described in Part 1 of this series pointed out the irony that while life needs the presence of water to operate, to get life started water must be periodically removed. The drying down of solutions is anathema to many solution chemists so this was a blind spot for those tackling the problem of the origin of life.
Also lying beyond Darwin’s 19th Century understanding was the science of complexity, specifically the generative and adaptive power of networks. Railway and steamship systems interconnected by telegraph communications networks were only just beginning to inform the future of control theory and automatic computation in the mid-Victorian period. Complex chemical networks with their feedback loops would only come to be understood in the 20th century during the period of the rise of cybernetics, computing and the Internet. As we shall see in a later article in this series, there is a central role such networks can play in the origin of life.
Proteins and other polymers participating in biology such as RNA and DNA all form through condensation reactions in which water must be a leaving group. Today, enzymes perform the function of removing water to form the ester and peptide bonds of polymers. As described in Part 1, before life there is only one mechanism able to consistently form long enough chains of polymers to express catalytic activity: the removal of water through dehydration. Drying can only occur at the interface between water, mineral surfaces and the atmosphere. This directs us back to Darwin’s warm little pond but updates it to a hot little cycling pool subject to cycles of drying and refilling to form increasingly complex polymers. On the volcanic land masses of the ancient Earth formed by upwelling magma, distilled fresh water evaporated from the early oceans would have fallen coming into contact with active volcanic systems. Hot springs periodically refilling a system of interconnected pools are and ideal “chemical engine” to transform a simple primordial chemical soup into colonies of living microbial communities.
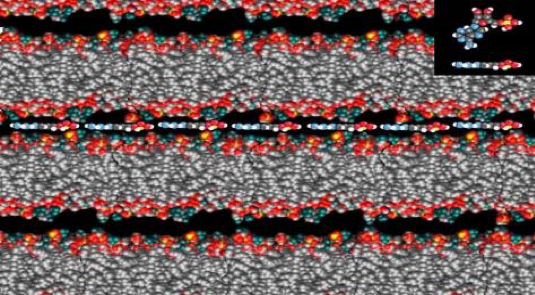

Figure 7. Computer graphic of lipid bilayer membranes sandwiching together the building blocks of RNA (inset).
If we imagine ourselves bending down to study one of these pools, we will notice that most of the chemical action occurs at its edge where water laps up on mineral surfaces. In ponds, lakes, rivers and the sea shore today, the boundary between water, mineral and air are the richest interfaces for diverse life and this suggests that this boundary may been where life originally began. As Dave Deamer did in that pool in distant Kamchatka, if you add mixtures of membranes, like those found in your favorite bar of soap, a sort of “bathtub ring” will naturally form at this boundary. If you took out a microscope and studied the dried films in that ring, you would see numerous layers sandwiching together smaller molecules from the bathtub, and you may detect some of your own DNA, RNA or proteins! If the building blocks of these important biopolymers were trapped there, they would be organized and stitched together as water left through the membranous layers (figure 7).
Our group at UC Santa Cruz and several others have formed these polymers in the laboratory in this way. More recently we have observed the same phenomenon in natural hot spring analogs for the early Earth. We can produce RNA-like polymers in lengths of hundreds of chain units, long enough to express functions: either folding to create a tool to do a job such as catalyzing a reaction, or to store and express information permitting more polymers to be assembled from a blueprint. Other groups have repeated our experiment and also demonstrated the formation of peptides, short sequences of amino acids which life produces for the cell’s toolkit: proteins.
The final step in our scenario occurs when the hot little cycling pool refills with water. The outer layers of the lipid “bathtub ring” bud off trillions of small compartments. Contained within some of them will be some of the polymers randomly assembled within the layers. We call these compartments “protocells.” Each of these protocells is a natural experiment in combinatorial chemistry. In the first cycles, some “pop” and some “do not” kick-starting the first form of selection. In summary, these cycling pools indeed possess the power to form Darwin’s protein compounds and other polymers and subject them to more complex changes.
Considering life’s origin through the lens of computer science
While Darwin was drafting On the Origin of Species, his contemporary Charles Babbage was designing a new breakthrough in Victorian technology: automatic mechanical computation. By the early 20th Century, devices derivative of Babbage’s unbuilt difference engine were transforming economics, science and culture. Alan Turing’s 1930s work on a theoretical universal computing architecture famously involved the reading and writing of symbols to a tape. The 1953 discovery of the structure of DNA suggested parallels to the punched paper tapes and cards of early electronic computers.
But how does the formation and combinatorial selection of polymers lead to a system in which life, a sort of program run through chemistry, can write, run, test and rewrite itself all without a programmer? From the perspective of computer science, a living cell resembles an elaborate chemical computer, but can a chemical computer emerge spontaneously on a sterile yet habitable planet like the early Earth? The answer is obviously yes, because life did begin, but the process by which this occurred remains a fundamental problem in biology. We will argue that by analogy to the development of computers, the hardware of the first, or indeed any form of life, is operated by: organic molecules organized into supramolecular structures capable of capturing free energy available in the environment and using it to drive polymerization and growth. We propose that the first programs of life spontaneously developed when initially random systems of polymers underwent cycles of selection and amplification to express functions within a system of self-assembling hardware built up from protocells.
But you might ask: how it is possible for programs to emerge in the absence of a programmer? Figure 8 illustrates how this can work by referring to early computers that used holes punched in a paper tape as a way to code binary bits. The goal is to generate a program that will turn on a series of light emitting diodes (LEDs) in a front panel attached to the computer. At the outset, the tape puncher is linked to a random number generator. The punched tapes are then read one at a time into a simple computer which uses electrical energy to read each tape and then executes the purely random instructions through a primitive processor. Most programs “crash” and no LEDs are switched on. However, by chance, a rare random program such as one labeled “A” in the example happens to light up at least one LED. This program gets to “play again” so is automatically selected and fed back into the puncher. The puncher replicates the program “A” and punches it again three times. For each new copy of “A” it also adds new random instructions “B”, “C”, “D” to the end of the tape. The next cycle runs the new programs A-B, A-C, A-D, and if, for example, A-C turns on another LED, it is selected to go forward into a new generation of programs. Over time a group of programs (A-C-F, A-C-F-etc.) will emerge which turn on more LEDs on the front panel. This method of programming is inefficient but given enough time it will create a program that switches on all of the LEDs.
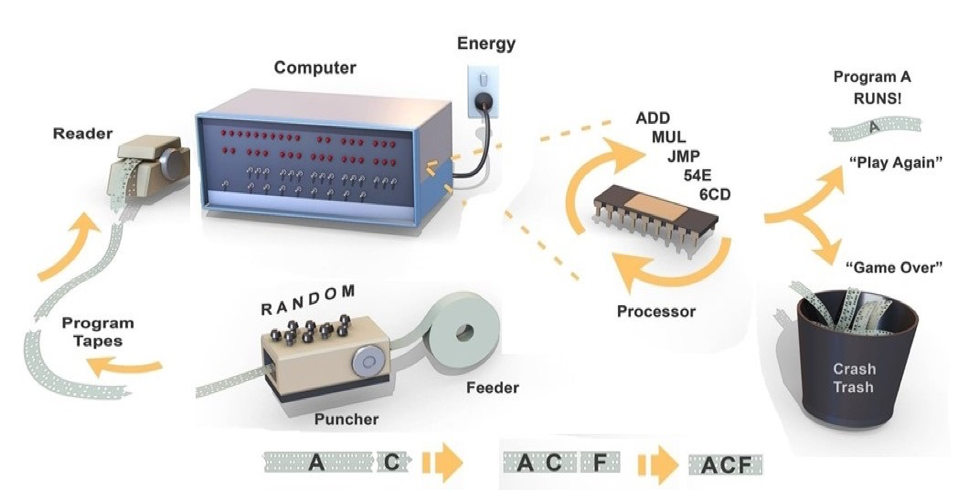
Figure 8. Metaphor for how a computer program can be developed without a programmer.
Coding a chemical boot-strap
To apply more chemical terms to the above auto-programming example, 1) a source of energy (an electrical current) drives a process that punches random holes in a tape. At some point, one set of holes happens to cause an LED to light up and that set is 2) selected, 3) reproduced and 4) amplified, within a population of continuing random hole punching until another chance sequence of holes causes a second LED to light up, and those sets are ligated together and so on until all of the LEDs are operating. A fifth, often overlooked intermediate step is essential, which is a simple set of instructions operating a feedback loop, a subroutine that instructs the system to save, incorporate and amplify any chance program that happens to turn on an LED.
These five steps, equivalent to a simple operating system’s boot-strap, iterated an enormous number of times on large systems of polymers can lead to an origin of life. The energy source is sunlight, the paper tapes are polymers such as nucleic acids or proteins, and the holes are chance patterns in monomer sequences. The programs are collections of those polymers that undergo selection and amplification if they happen to have functions that increase survivability of those same polymers over time. The collections are contained within membrane-bounded compartments called protocells.
The metaphor of a self-writing computer code leads us to reframe the origin of life in the form of a testable hypothesis that incorporates the biophysics of self-assembly, the chemistry of polymerization and the geophysical properties associated with volcanic land masses on the early Earth. Although this reframing provides a new perspective for life’s origin, it also raises many questions. First, we must define what we mean by an operating protocell. The sole measure of operability of a microbial cell is that encapsulated systems of catalytic polymers use nutrients and energy to undergo growth and reproduction through polymerization. Akin to the punched paper tapes, the system we propose generates immense numbers of microscopic protocells, each different in composition from all the rest, and these protocell populations must then undergo a primitive version of Darwinian selection at the molecular and supramolecular level.
This brings us to the set of questions we are exploring here:
- How can the earliest protocells composed of primitive, self-assembled membranes encapsulate cargoes of random polymers?
- How can chemical “programs” spontaneously emerge by selection and begin to operate?
- What environmental stresses provide selective factors, and how can programs be re-written so that protocells can survive these factors and propagate sets of polymers into subsequent generations?
- Are there prebiotically plausible natural environments in which protocells can emerge and then undergo evolution through natural selection before the emergence of life?
- How can such populations of primitive protocells eventually evolve into the self-sustaining and self-reproducing state that defines living systems?
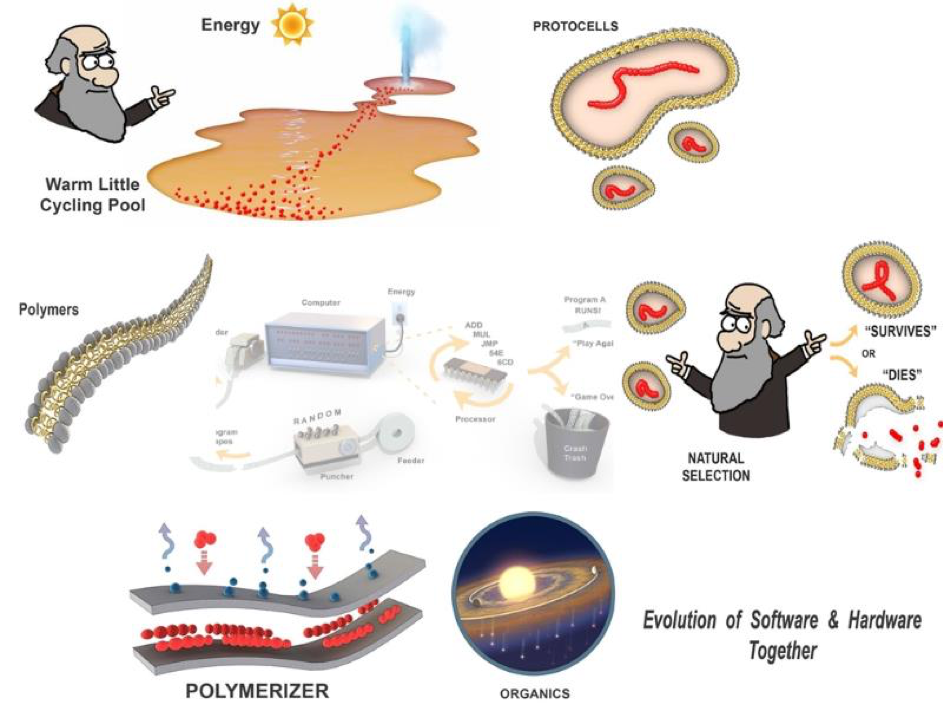
Figure 9. A natural setting and equivalent components for a self-programming chemical computer boot-strap.
But where can we find a natural setting that behaves like our primitive self-programming computer setup? Why, back in Darwin’s warm little pond, but updated to the hot little cycling pool shown in figure 9. The “punched paper tapes” of life are polymers built up out of monomers such as amino and nucleic acids by our “polymerizer”, layers of lipid in our drying little pool which stitch together these monomers. The polymers have the active (structural or catalytic) functions to be “read” and “run” in our little pool. They are packaged as sets of random “instructions” within membranous protocells. Each protocell is “executed” by our program tester, represented by our cartoon Charles Darwin. He applies simple logic called Natural Selection and each protocell program either survives his test or succumbs to disruption losing its contents and “dies”. Surviving protocells return with their polymer cargoes for another round in the polymerizer. In this manner, a completely physical process can create a system that is able to select functions out of the background of random sequences. In this example, those polymers which keep their surrounding protocell from popping are the first function selected for. In this way evolution through what we refer to as combinatorial selection can get underway before life. In other words, a simple form of evolution is the necessary boot-strap that precedes life. Any hypothesis is just “hand waving” unless it can be subject to rigorous testing and in the next part, we will update you on the laboratory and field science to test (or falsify) this scenario.
Part 3. The Quest to Test (or Falsify) the Hot Spring Hypothesis
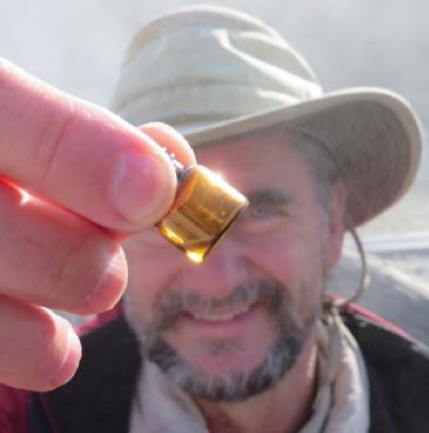

Figure 10. The author as a sort of “Indiana Jones” of Astrobiology in a quest to synthesize protocells in the wild, polymerizing RNA by wet-dry cycling acidic spring waters at Hells Gate, Rotorua, New Zealand, June 2018. Photo: Kathleen Campbell.
The newly emerging interdisciplinary field of Astrobiology involves adventuresome science worthy of an Indiana Jones film (figure 10). From rovers on Mars to telescopes high on mountaintops and in space searching for distant habitable worlds, the search for life in the universe takes human inquiry to truly new frontiers. The origin of life plays a central role in the field of Astrobiology yet its main workers are chemists, who for decades have remained cloistered in their clean laboratory environments. In the past few years, however, a series of collaborations between chemists, computer scientists, geologists and others has resulted in a major move beyond comfortable campus science out into the wild. Part 3 will take us from the lab to the field to track some of our work to test, or falsify our scenario at the whims of Mother Nature.
In Search of the Goldilocks setting for an origin of life
Geologists are now in general agreement that oceans could have been present on the Earth as far back as 4.3 billion years ago. The cooling process of the Earth’s mantle led to vast volcanic eruptions which continued for billions of years and would have produced the first large land masses, well before the rise of continents riding on their tectonic plates. While well over 90% of the Earth’s liquid water would have been present in the salty ocean, while evaporation from the ocean and subsequent rainfall would have delivered fresh water to these land masses. On volcanic land masses today, rainwater finds its way into rock cracks and cavities in active volcanic fields, is heated and driven back up to the surface, often with clockwork regularity like the original Geyser in Iceland or the famous Old Faithful at Yellowstone National Park in the USA. Such a hydrothermal field on an ancient volcanic landscape over four billion years ago would have provided an ideal, or “Goldilocks” setting for life to begin. In the well-known fairytale, a young girl named Goldilocks was walking in the woods and discovered the house of three bears and decided to try their unattended bowls of porridge until she found one that was “just right” (not too hot, not too cold).

Figure 11. Goldilocks in search of the perfect porridge. Image: L Leslie Brooke.
Let us make up our own version of the tale with the young Goldilocks working in the kitchen to make “just right” bowls of porridge for the three bears (figure 11). She would have first heated some water on a stove until it was “not too hot, and not too cold”. Next, she would have found a small bowl, adding the water to the bowl together with all the right ingredients and stirred them together. She would have then started sampling the porridge, one spoon at a time, as she sprinkled in flavorings, ultimately declaring the moment when the whole bowl was worth the eating. She might have then made a bigger pot and doled out porridge into three bowls of different sizes and set them out for the three bears’ return home.
The Goldilocks metaphor is actually quite apt for understanding our optimal “primordial porridge” at the start of life. Fresh water upwelling from the stove of our volcanic spring finds its way into a small bowl-like pool. The water is just hot enough for the key chemical reactions to happen but not too hot. Falling in from the sky and flowing in from the surrounding landscape and deep from within the hot spring itself are our starting ingredients. Those ingredients are stirred together and form spoonfuls called protocells. Each protocell is tested like a natural experiment and selected to form more protocells. Our protocell porridge gets tastier and tastier as it uses more ingredients. Bits of this porridge find their way into other bowl-like pools and a whole cookbook of porridges of different sorts emerges across the table of our landscape. Mixing recipes provides more elaborate porridges until one day these porridges have written their own unique recipes so they can cook themselves up from available ingredients and energy.
A century of origins of life experimentation and speculations
For more than a century after Darwin’s 1871 “warm little pond” insight science has been trying to make this original porridge by trying many recipes. In the 1920s Alexander Oparin and J. B. S. Haldane proposed that a concentrated enough “primordial soup” perhaps composed of lumps of protein they called coacervates could form a medium for an abiogenesis. In the 1960s, Graham Cairns-Smith proposed that it was the clay edges of pool, the bowls themselves, which could promote the formation of polymers.
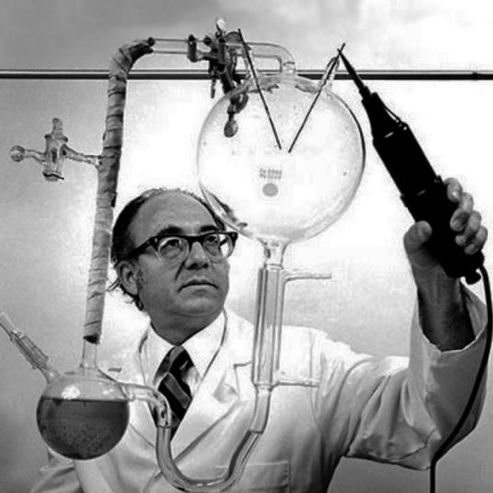

Figure 12. Stanley Miller with the famous spark chamber setup of their 1952 experiments.
But it was the experiments of Harold Urey and Stanley Miller in the early 1950s (Miller, 1953) in which they provided an electrical spark discharged into a vessel containing a gas they supposed represented the early Earth’s atmosphere (figure 12). These discharges produced black deposits on the sides of the vessel which contained important organic compounds, notably dozens of kinds of amino acids, the building blocks of proteins. Their discovery was announced with great fanfare in the same year as Watson and Crick published the structure of DNA, and this work launched the field of the origin of life as an experimental science.
The 1977 discovery by the submersible Alvin of deep oceanic hydrothermal vents, devoid of sunlight but housing a rich chemical-eating ecosystem from microbes up to meter long tube worms and numerous fishes, was put forth as a possible place life could have started (Corliss et al. 1981, Russell et al. 2014). However, decades of experiments in laboratory simulations of vent environments have failed to produce more than trace amounts of the amino acids or any the other building blocks of biopolymers or membranes. Proponents of the vent hypothesis have put forth mineral compartments or gels as early stand-ins for protocell compartments, as membranes cannot form stable enclosures in sea water (Milshteyn et al. 2018). However, the thermodynamic barrier of the “water problem” described in Part 1 will simply not go away in an environment under a thousand meters of ocean. Yet another challenge is that if a form of life started at a deep-sea vent there would be no way out for that life. Surrounding the vent is a vast bulk of deep, cold and dilute water. Without vent chemicals to sustain it, fragile proto-life straying from the immediate hot spot would starve. There would also be no way for vent life to develop photosynthesis as an alternative food source in the darkness of the deep ocean before it found its way to the ocean’s surface or very shallow coastal sub-surface. For such an adaptation it would have to have access to both hydrothermal chemical energy and sunlight for an extended period of time to adapt to entirely depend upon this new form of energy capture. In other words, to develop photosynthesis, early life would have to evolve where it had consistent access to both energy sources… in the margins of hot springs on land!
Returning to Darwin’s warm little ponds (or cycling hot little pools)
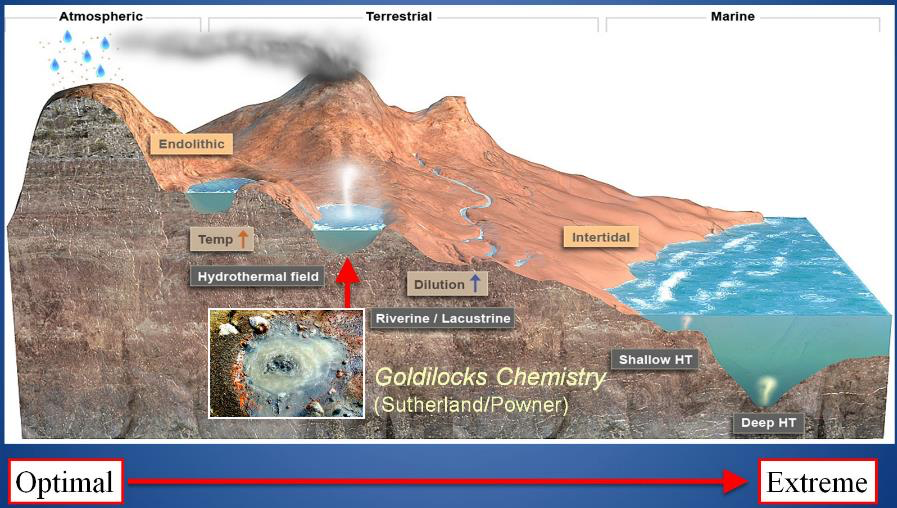
Figure 13. Where we might find pools on land able to support a “Goldilocks Chemistry”. Image: Bruce Damer and Ryan Norkus.
As a result of these seemingly insurmountable barriers to an origin and early evolution of life in the deep-sea vent environment, workers in the origin of life field have returned their focus to pools on land, seeking a “Goldilocks chemistry” called for by (Powner and Sutherland, 2011) in pools that were not too hot, not too cold and where the porridge (or perhaps the primordial soup) is just right. Originating in optimal fresh water cycling pools and fed by extraterrestrial and hot spring ingredients, proto-life would have many different pools to distribute and adapt to. Pool variations in pH, temperature, dissolved solutes, concentrations of organic compounds and cycling rates provide a “combinatorial landscape” for protocells and their polymers to form and mix under differing conditions. Figure 13 illustrates how protocells and their progenote aggregates might start in one Goldilocks pool and distribute downhill (or be blown by wind as a dried film) into many other aqueous bodies such as dilute streams and lakes, in watery cracks in rocks, and then on to the more extreme salty seashore. For life to escape one Goldilocks environment (such as a hot little cycling and chemically well-supplied pool) to a cooler and dilute non-cycling “extreme” environment like an outflow channel, it must adapt to capture sunlight to make its own food. This pattern can still be seen today at Yellowstone in the outflow channels of sinter mount hot springs where chemical feeding microbes yield to orange and then green microbial mat communities living on sunlight in the outflow channel (figure 14).
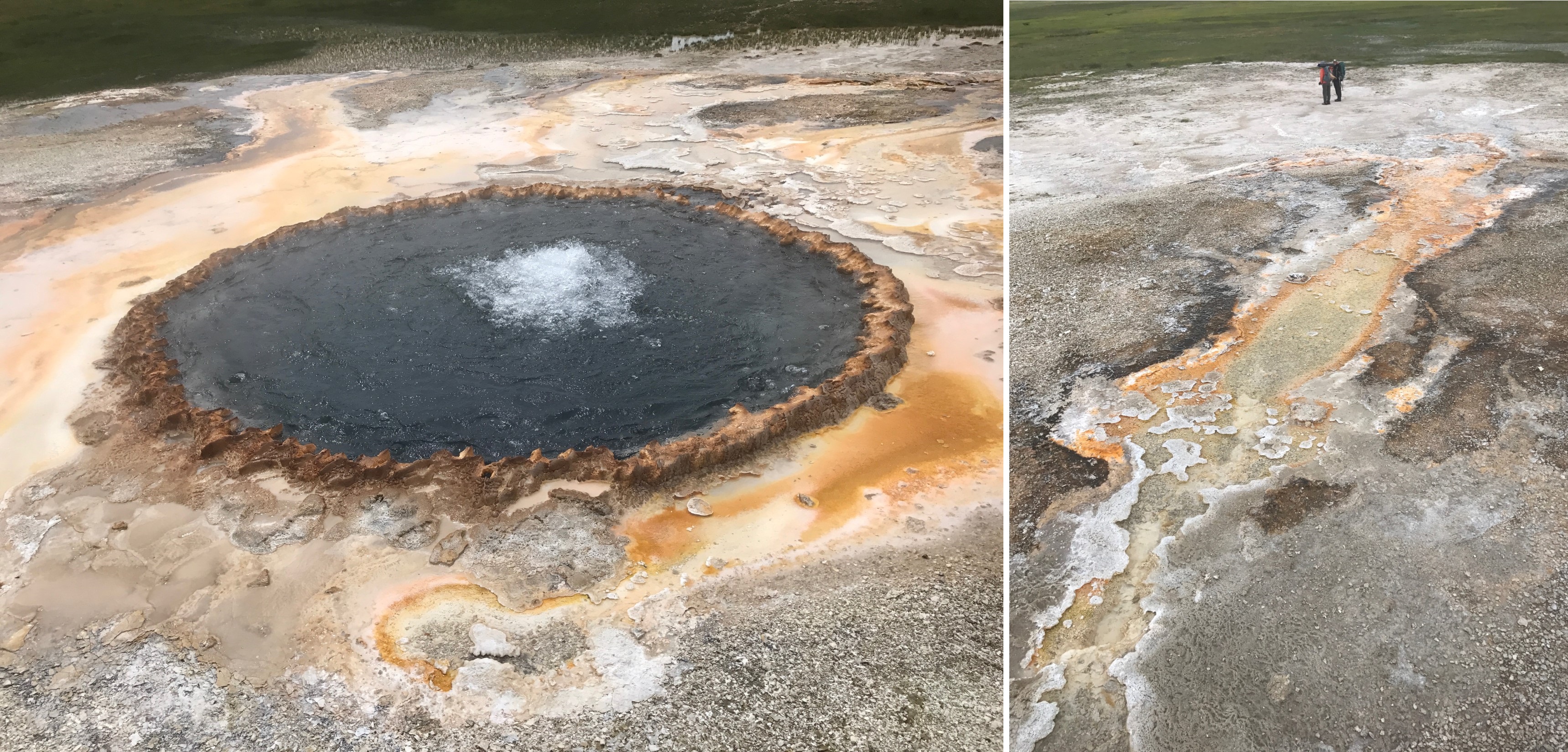

Figure 14. Left: sinter mound at Yellowstone National Park surrounded by microbes feeding on hot spring chemicals. Right: outflow channel with microbial mat communities adapted to high temperature photosynthesis and lower temperature green photosynthesizing plants in the distance.
Photo-capture could have begun with protocells utilizing a naturally occurring pigment such as polycyclic aromatic hydrocarbons which are a common component of meteoritic in-fall, but even a primitive form of biological photosynthesis is so complex that it argues that the transition to life must have occurred to enable it. Therefore, in our model the transition to living cellular communities with genetic materials and mechanisms to code for the expression of enzymes must have occurred at the boundary where hot spring waters cool and dilute. So, by this logic, perhaps we can predict the exact location on this landscape by which protocells must become early living cells. As the boundary of what is “extreme” for early life is pushed forward (or downhill) early microbial communities will encounter brackish water, first in drying pools and then in tidal estuaries. Salty water is highly stressful on membranes, especially prebiotically plausible, very flimsy ones such as those built out of meteoritic fatty acids. So active membrane transport with energy driven pores must have evolved to permit life to gain access to the marine environment. The open oceans or deep hydrothermal vents would be a still much later adaptation. Now that we have a view of the landscape on which life may have originated, we can turn to experiments to replicate their conditions in the lab and out in the field and test the hot spring hypothesis for life’s beginnings.
Wet-dry cycling in the laboratory and the field
For decades experimentalists were frustrated by ending up in many chemical dead-ends. A promising reaction could be demonstrated, but only in simple laboratory conditions and the reaction would eventually run down and the products ultimately degrade in the presence of water. A hint of a solution to the water problem of the origin of life presented in Part 1 was first presented by researchers who demonstrated that amino acids could form peptide bonds in hot, drying conditions (Sawai et al. 1975). It was Prof. David Deamer who demonstrated nucleic acid polymerization in the most obvious way: by the removal of water through dehydration.
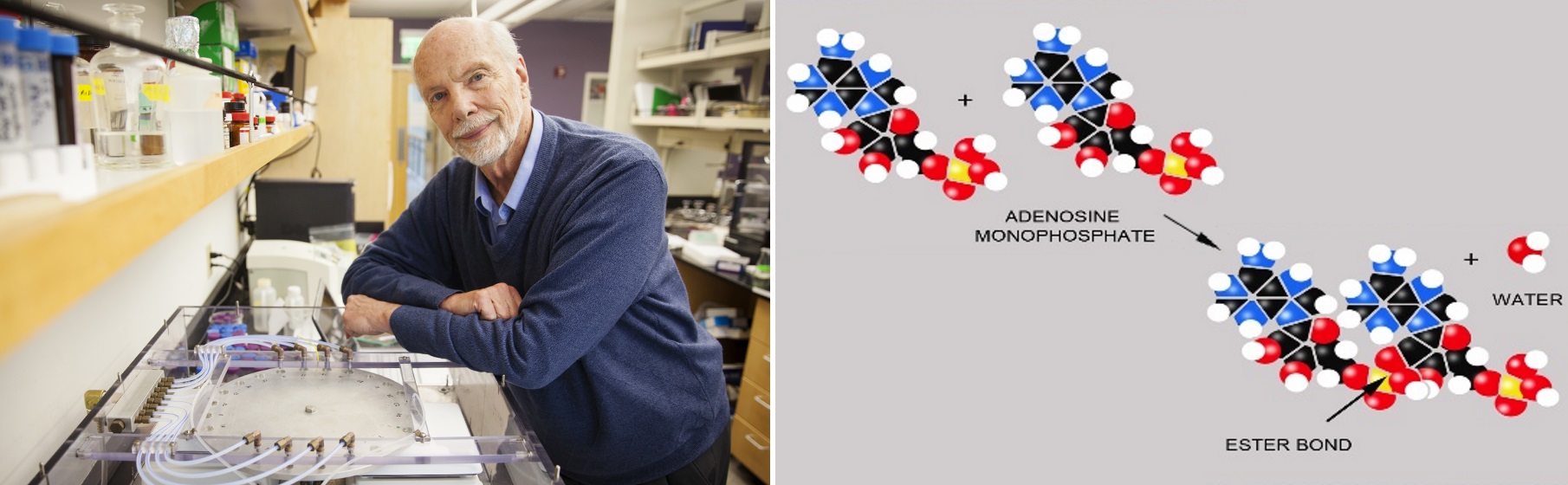

Figure 15. Left: David Deamer at his lab at UC Santa Cruz with the wet-dry cycling simulation chamber. Right: The formation of an ester bond between RNA monomers of adenosine monophosphate (AMP) by making water a leaving group through dehydration.
Deamer and colleagues showed that two of the building blocks of RNA, AMP and UMP could also stitch together to form long polymers of RNA, in some cases in excess of 100 units. This is significant as at this length RNA strands could perform jobs as catalysts or the storage of short pieces of information. This supported a prevailing view known as “the RNA World” that RNA arose first, as a double duty polymer, performing the information storing role of DNA and the functional roles normally carried out by proteins. To “grow” long strands of RNA, Dave and his team had added the “secret ingredient” of lipid to provide an organizing matrix for the RNA building blocks. Dave and his team built a simulation chamber (figure 15) in which mixtures of lipid, AMP, UMP and slightly acidic water to promote the reaction, were subjected to cycles of hydration and dehydration. Early Earth atmospheric conditions were simulated by removing oxygen, and mixtures were heated to up to 90C to match hydrothermal field environments and increase the rate of the polymerization reactions.
As the vials dried down, lipid layers would form on the sides and bottom of the vials, Lipids are the class of molecules which today form all cell membranes. Simple forms of lipids have been observed to be formed in space and delivered to Earth on asteroids and dust particles. The layers sandwiched two of the building blocks of RNA (AMP and UMP) into a 2D layered space which oriented and pre-stacked these monomers so that when water became a “leaving group” (figure 15, right) through dehydration, the RNA polymer was formed from individual building blocks (Rajamani et al. 2008, Deamer et al. 2019).
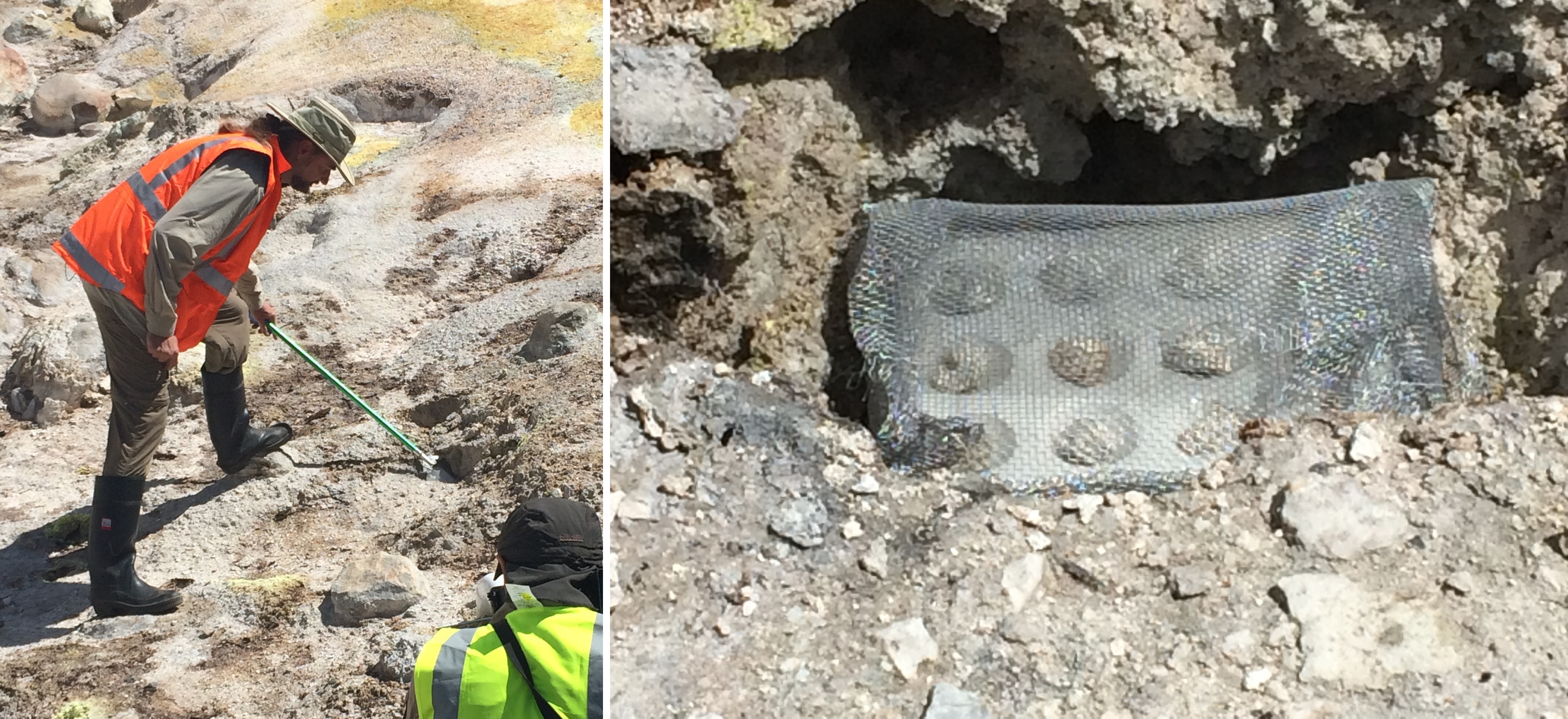

Figure 16. Left: placing a sample into a fumarole vent in the hydrothermal field at Bumpass Hell, Mt Lassen Volcanic Park, California. Right: mineral chips set into well plate in fumarole vent.
Dave was a great believer in leaving the laboratory to attempt to try experiments in the messy, complex conditions at active volcanic hydrothermal fields. After two trips to the Kamchatka peninsula in the far east of we journeyed to the steaming fumarole vents of Bumpass Hell, at Mt. Lassen Volcanic Park in Northern California to try to use the humid vent gases to form RNA in drying-down glass wells (figure 16, left). After many failures, on our August 2016 trip we were able to detect a small amount of RNA-like polymer on mineral chips exposed to wafts of these moist gases (figure 16, right).
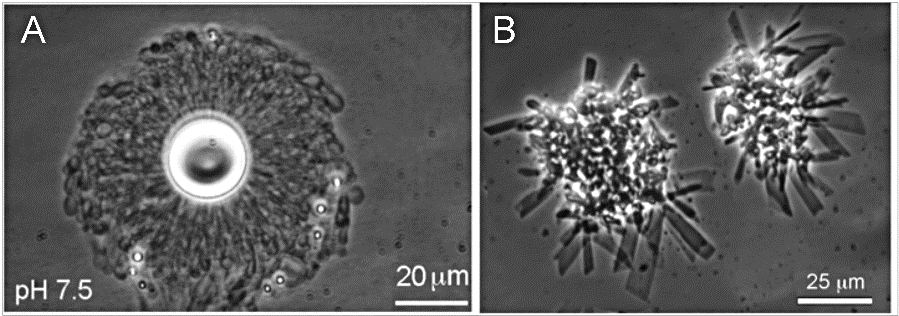
Figure 17. Comparison of vesicle assembly in Yellowstone hydrothermal water (A-left) and seawater (B-right). The divalent cation content of seawater causes the fatty acid to form insoluble crystals rather than membranes. Photo: Milshteyn et al. 2018.
We then decided we needed to try our experiments using water from active hot spring pools so visited to Yellowstone National Park with colleagues in June of 2017 to mix acidic and alkaline hydrothermal waters in a number of vials containing dried lipid and other solutions. Holding up a vial the milky liquid indicated we were forming trillions of vesicle compartments. Under a microscope at the lab at UCSC, stable vesicles were indeed formed with different samples of Yellowstone water (figure 17, left) but when we added filtered sea water they collapsed, forming crystals (right). The vesicles formed in the hot spring waters were numerous and able to encapsulate DNA after being wet-dry cycled (Milshteyn et al. 2018). So, we could “cook protocells” in a Goldilocks pool (but not a salty one)! These results provided crucial support for a freshwater origin for life.
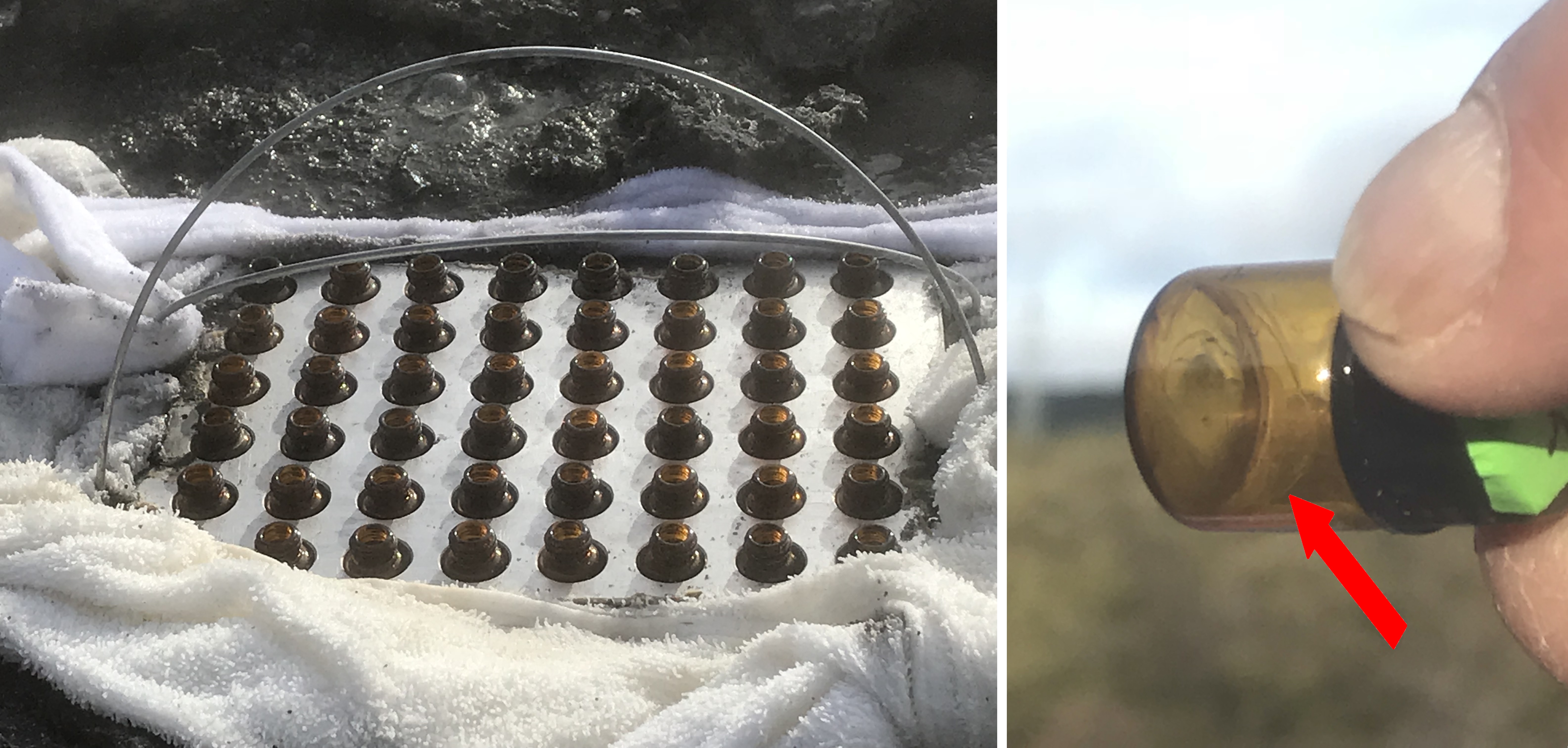

Figure 18. Left: vials in an aluminum well plate set into a 93C pool at Hells Gate, Rotorua, New Zealand. Right: layers of dried lipid visible in vial after a 15-minute drying cycle.
In June of 2018 we traveled to the Hells Gate Geothermal area, on the North Island of New Zealand and cycled vials of dried lipid together with AMP and UMP building blocks through four wet-dry cycles over four hours. The vials were sequentially filled with a small drop of hot spring water drawn from nearby pools – half with acidic (pH 2) water and half with slightly alkaline (pH 8) hot spring water. The vials were left open to the sulfur gases and high humidity of the environment, and set into an aluminum block directly in contact with a 93C bubbling pool (figure 18, left). Layered films of lipid clearly formed on the inside of the glass (figure 18, right) but were the monomers of RNA shuffling around between these layers and forming polymers?
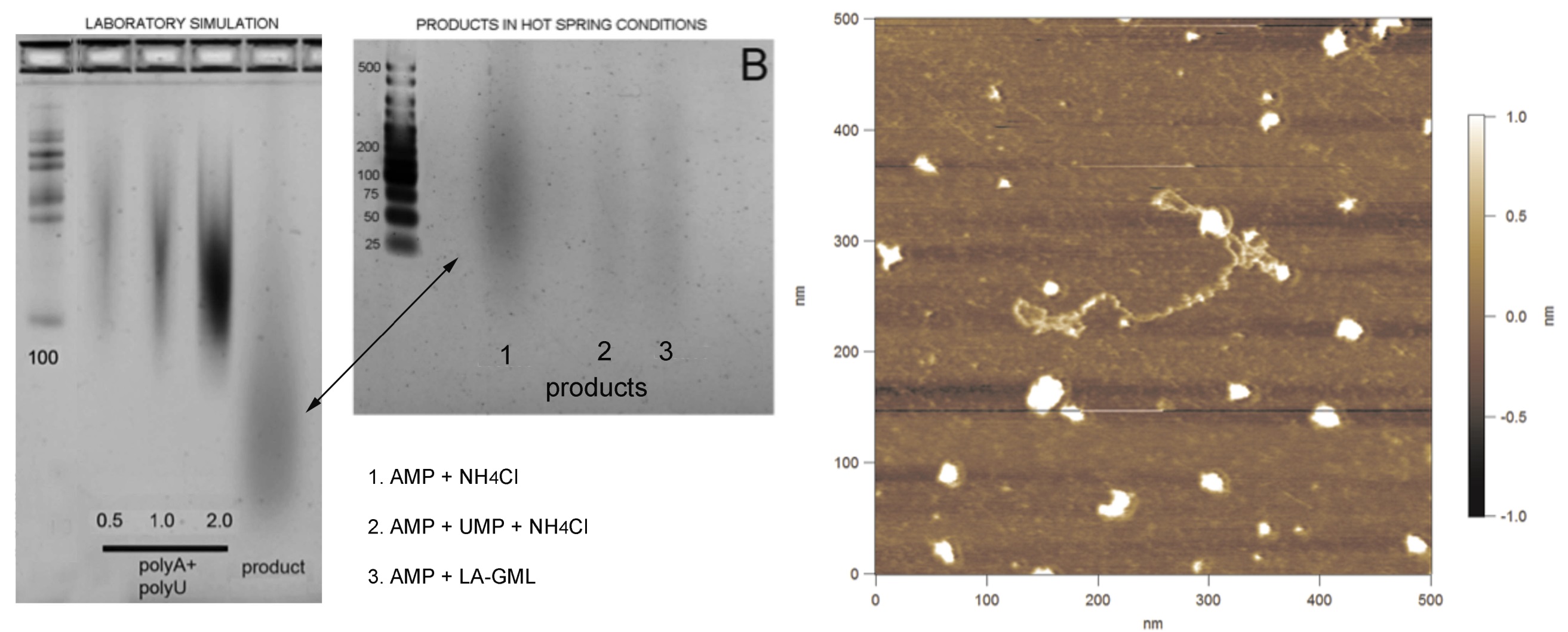
Figure 19. Left: comparison of polymer products from a laboratory simulation using 4 wet-dry cycles (left image) and field-based cycling using acidic hydrothermal water from the Hell’s Gate hydrothermal system near Rotorua, New Zealand (center image). Right: fibril imaged by atomic force microscopy of putative RNA-like polymer in the Rotorua sample. Image: Tue Hassenkam.
Back in the lab at UCSC we could finally compare our laboratory “simulations” with our hot spring field analog experiments. We used gel electrophoresis to separate RNA polymers by size. The samples are loaded in the top and when an electric current is applied, the polymers move through the gel with smaller polymers moving faster and further through the gel than longer one. The left lanes on each gel show bands of polymers of known length (a ‘ladder’) which we use to estimate the length experimental products. Figure 19 (left) shows three experimental control lanes (denoted polyA + polyU), where synthetic polymers were made from strings of AMP monomers (polyA) and UMP monomers (polyU). Using the ladder, we can see that these polymers are between 100-200 units long. The experimental lane on the right shows the polymer produced by cycling a mixture of AMP and UMP in the presence of a lipid (LPC). The product traveled further through the gel than the 100 unit standard, so we know it contains polymers 20-100 units long. Similar streaks in that range can be seen in the products synthesized by cycling AMP and UMP monomers in acidic water of the Hells Gate hot spring (figure 19, center image, connected by arrow). The image in figure 19 is a preliminary result, an image of a “fibril” that may be one of our RNA-like polymers imaged from the Rotorua samples by atomic force microscopy. So, for the first time, we may have been able to “grow protocells in the wild.” One caveat is that these experiments were still carried out in vials, protected from the diluting effects of an active pool. In our next fieldwork we plan to introduce dried powders of lipid, RNA monomers, and possibly also amino acids, directly in contact with mineral surfaces and water in hot spring pool settings. This work is well reported in (Deamer et al. 2019).
This laboratory and field work are promising and several university teams around the world are duplicating it and preparing a new generation of young investigators to don field gear and good boots to engage in more bold experiments. But can we say that we know for certain where and how life started? Definitely not, but perhaps we can state that we can now make more informed guesses. Science judges the value of any hypothesis by the testable predictions that it makes and as those predictions turn into evidence they are weighed. Parts of a hypothesis that don’t stand up to testing must be discarded early to make way for more testable modifications to the hypothesis. At some point a hypothesis may accumulate a weight of so much evidence that is graduates to becoming a theory. The hot spring hypothesis is nowhere near its own graduation date but perhaps we could say it is looking like it has a strong prospect of making it through its freshman year! In the next and final essay, we will leap forward from our promising platform of experimental foundations and make some new informed guesses about how this scenario for the origin of life can be linked to its subsequent evolution, and speculate about some implications carried for the Extended Evolutionary Synthesis.
Part 4. The Progenote, Chickens, Eggs and the Extended Evolutionary Synthesis
First inspired as a teenager by the question “how did life begin?” I reframed it as a new question: “how can you make a copy of a machine without employing a bigger machine?” After decades of research and computational work, a thought experiment occurred to me on the morning of December 30, 2013 in which a network of interacting little machines (protocells) cycled and collaborated to collectively become a bigger, more complex machine (a living microbial community). This thought experiment was sketched in figure 20 (left) and became the basis for the “coupled phases” scenario for an origin of cellular life (right) published by Damer and Deamer (2015, 2019) and described in the first essay in this series. Briefly, at the bottom of a hot spring pool during its drying phase, lipid layers form on its mineral surface which organize and link together concentrating monomers into random sequences of polymers. These polymers are then encapsulated into vesicles budding off when water flows back into the pool, forming large numbers of protocells, which contain random sets of polymers. Each protocell is an experiment in combinatorial chemistry that can select for functional polymers on the first few steps toward life. As it turned out, however, the scenario which emerged from my thought experiment was not yet complete.
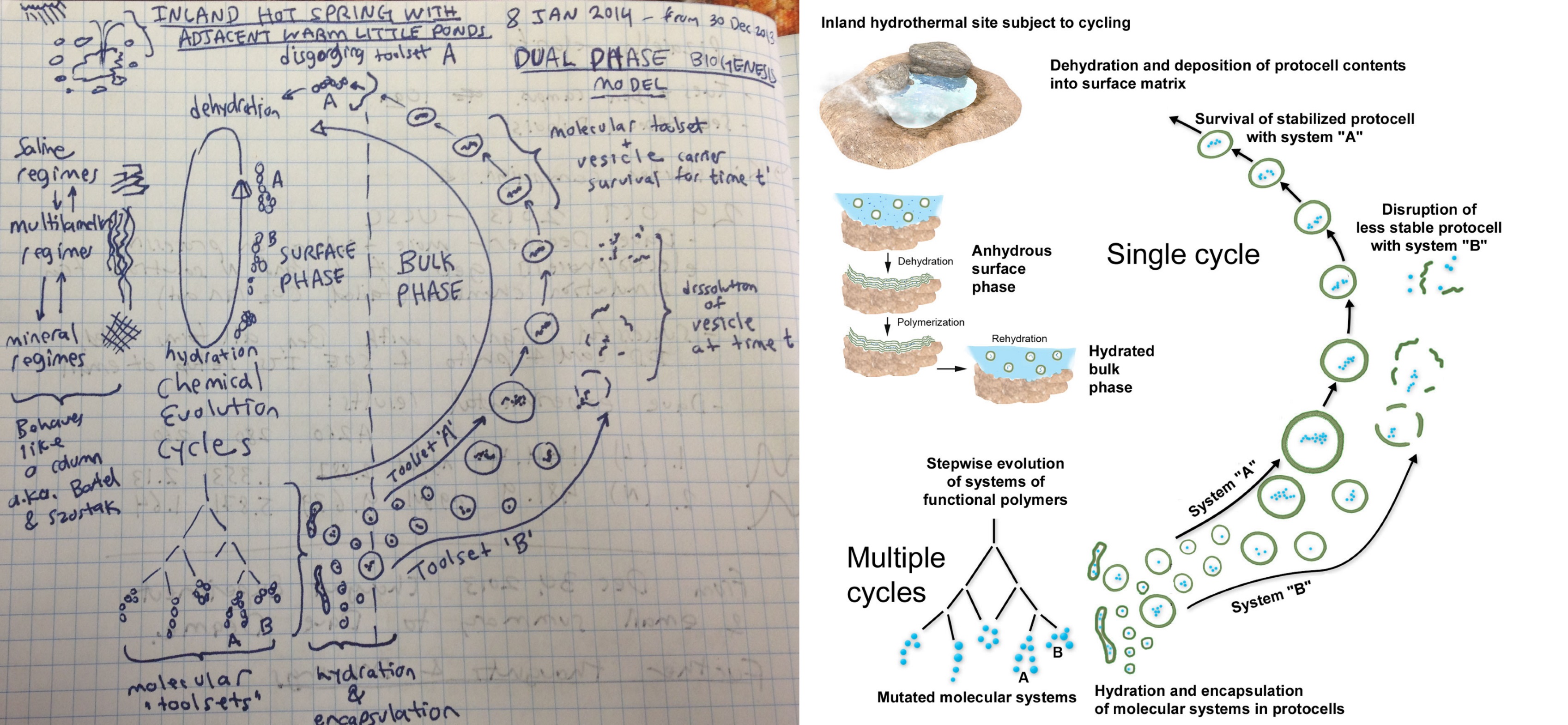
Figure 20. Left: the sketch made by the author of the scenario a population of cycling protocells undergoing combinatorial evolution derived from a thought experiment on December 30, 2013. Right: its development into the scenario in the “coupled phases” hypothesis (from Damer and Deamer, 2015).
During a 2016 field trip to Western Australia, another realization occurred that between the dry (layered) state of the lipid enclosures and the wet (free floating protocell vesicles) there must be an intermediate, moist phase in which stable, surviving protocells aggregate together at the bottom of a drying pool. I termed this intermediate form the “gel phase” as it would be a hydrogel composed of lipid membranes and water, and shared it with my UC Santa Cruz colleague David Deamer who was seated on the bus next to me. He commented that “it would be selected for” as the structure of the gel would act to protect its fragile member protocells. In other words, there can truly be “power in numbers.” It also occurred to me at the time that these aggregates would concentrate key components to support the first metabolic activities within protocells. In addition, the products of reactions in one protocell could diffuse through the gel to other protocells, setting up a network effect of interacting chemical circuits. The analogy I drew from was that the normally dry Australian desert we were driving through had recently experienced a rainfall, and its soils were moist and bursting with shared metabolic activity as evidenced by the splashes of green vegetation and color from roadside flowers. The drying cycle would arrive just as these same desert plants budded off their seeds – protective capsules to protect and distribute their DNA ready for the next wet cycle. This scene was an elegant depiction of the ubiquitous life cycle in the natural world, perhaps suggestive of a distant echo of a simpler three-way cycling system present at life’s origin.
The Progenote as a proto-niche
Months later, during visit to the University of Houston I met with Prof. George Fox, who had worked as a young research associate with Carl Woese. He suggested that the three way cycle we were proposing was suggestive of a pathway to the “progenote” that he and Carl had described (Woese and Fox, 1977; Woese 1998, 2002). I realized then that our gel phase aggregate was a candidate progenote, the unit of selection and combinatorial evolution that could cross the “chemical evolutionary chasm” to the first microbial communities. The concept of the progenote as a key player at the origin of evolution was described in Part 1 in this series. In this final essay we will take a deeper dive into the origin of life scenario, the progenote itself, explore connections with the Extended Evolutionary Synthesis (EES) and implications for this first major transition in evolution. Stepping back and taking the long view, we will seek a new understanding of the evolution of life on Earth from microbial communities to complex life such as algae, fungi, land plants and animals. We will conclude by proposing an answer to the classic question posed in the introduction: which came first, the chicken or the egg?
Searching for a link between the self-assembly of primitive aggregates of protocells and modern microbial mat communities, I discovered the work of John Odling-Smee, Kevin Laland and Marcus Feldman on Niche Construction Theory (NCT) (Odling-Smee et al., 2003). In later meetings with John and Kevin the term “proto-niche construction” was proposed as a way to describe the purely passive, self-assembled formation of these first protocell aggregates in which no active (enzyme-driven) work is being done by their members. The structure of the gel aggregate is built by entirely physical processes and source materials supplied from outside yet through many cycles, molecular evolution similar to that shown by Bartel and Szostak (1993) can occur and surmount selection hurdles. In this manner the passive proto-niche is the first step on a long, circuitous journey to become first active biological ecosystem. At each stage, energy-driven but initially passive processes of self-assembly are taken over by active processes of interacting polymers. Early evolution would operate almost entirely through mechanisms of group selection of progenote populations because individual protocells cannot express much in terms of function or survive for long on their own in the dilute pools of the early Earth. Living cells require a substantial degree of crowding of highly tuned interacting polymers together with other smaller molecules. The first protocells can only contain a few polymers expressing simple functions such as, for example, the stabilization of the surrounding membrane. They are far too dilute to be self-maintaining and also contain many random inactive molecules, existing in a sort of “garbage bag world” each filled with dirty water as described by Freeman Dyson (1999). Metabolic activity only becomes possible during drying down, when concentrated materials crowd into the aggregating protocells.
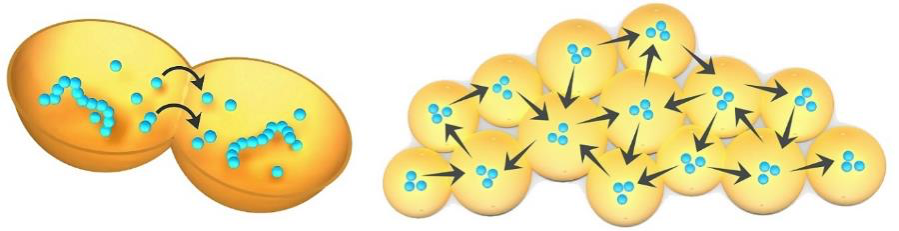
Figure 21. Graphical representation of diffusion of the products of one protocell into another and throughout an aggregate of protocells.
Despite the initial lack of internally driven active processes within member protocells, their aggregation still alters the local environment providing beneficial properties for its members. It is in this respect that the process resembles niche construction. Examples of these properties include physical protection from sheer forces, and mediation of pH and temperature. The membranous walls of the aggregated protocells would also help to maintain a collective transmembrane solute balance, thereby protecting all its member protocells against osmotic stress. Lastly, diffusion pathways within the aggregate can subject its protocell members to group selection. Figure 21 (left) illustrates how the metabolic products from one protocell can diffuse across adjoining membranes into the interior of another. With increasing metabolic activity, such products, and whole complexes of functional innovations can diffuse throughout the aggregate (right), benefiting the entire population. This property is a primitive form of a network effect which evolves into the consortium behavior and horizonal gene sharing characteristic of all microbial communities, suggesting that these modalities may have been central to their origin (Woese 2002).
The progenote has an additional powerful property: it can be distributed, both as a floating mat, washed downstream into other pools, rivers or lakes, or as a dried film carried aloft by wind to more distant aqueous settings. Microbial communities today use both of these mechanisms to spread. The dried phase of the progenotes would act as a preservation agent similar to seeds and spores. This is important as the longer polymers are exposed to water the more they will degrade through broken linkages due to hydrolysis. Therefore, before the existence of active enzymes able form or repair polymers, they must cycle through a drying phase. As progenote populations distribute across the landscape, many will fall into colder, more dilute settings such as streams or lakes. Most will cease to grow and simply dissociate as their rickety and poorly optimized chemical networks run out of building blocks and energy. Subject to such stresses, a rare few will develop robust innovations to adapt to these conditions such as the selection of pores, metabolic cycles, catalysts and the replication of informational and translational polymers. A key early innovation would be a mechanism to capture sunlight and utilizing this energy source to begin to synthesize needed building blocks. With photoautotrophy comes increasing distribution and independence from localized “Goldilocks” chemical conditions (Powner and Sutherland, 2011).
Within the aggregate of the progenote, its member protocells are engaged in the collective process of evolving the relationship between genotype and phenotype (Arnoldt et al., 2015). As individual protocells gradually accumulate and internalize proto-genomic functions, they will develop more autonomous phenotypic expression. A new evolutionary phase of the progenote will begin when member protocells can employ a primitive translation system to enzymatically replicate encapsulated genomic and functional polymers and pass them on to subsequent protocells without depending on dehydration synthesis. The combination of the controlled duplication of protocell contents with the division of protocell compartments will mark the last stage in the crossing of the evolutionary chasm to living cells. The growth and regulation of traffic across protocell boundaries and entire aggregates will be key to this transition as passive proto-niches become active ecological niches.
Crossing the Darwinian threshold, the progenote emerges as the first microbial mat community. The resulting speciation and specialization of increasingly well adapted cells will lead to microbial colonies becoming robust to steeper gradients of environmental pressures. These colonies can then adapt to more extreme environments on land and finally to the sea shore. Salty conditions necessitate active membrane transport to remove dangerous divalent cations in the much more extreme tidal marine environment. Eventually individual protocells become advanced enough to survive and reproduce independent of their source community, passing another important milestone in the evolution of life. Wider distribution and direct competition between communities and free-living cells extrude the many tangled evolutionary branches (or roots?) of early life along the long winding roads to LUCA, a last universal common ancestor.
The chasm-crossing progenote is a potent playground for creative thought experiments. However, caution must be exercised when developing scenarios, as evolution is not goal-setting and can only discover new molecular innovations by overcoming selection barriers. Plausible factors in the environment must therefore be considered. For experimentalists contemplating crossing this chasm in the laboratory, in effect creating an artificial abiogenesis, an additional caution is that this goal may be inaccessible. The early Earth had millions of years and innumerable cycling pools to undergo countless starts and failures, and a wide network of innovation-sharing across a landscape. However, the first few steps which increase the complexity of protocells beyond simple stabilization effects to primitive pores or metabolic catalysts may be experimentally discoverable, providing new insights about how life could have begun on Earth and where life might arise on other habitable worlds.
Looking through the long lens: relating life’s origins with its subsequent evolution
Leslie Orgel (1968) proposed that all living systems follow a principle of continuity in which new systems or structures are all built upon extensions to existing forms. In a computer’s operating system, the central kernel process can only be replaced by an entirely reworked code by the most skilled engineers. The engineer of evolution is much more conservative and cannot apply planning or intelligence so typically can only make minor selective alterations to existing kernels. Therefore, we might expect that the patterns we see in living systems today and throughout their evolution were established early, perhaps as early as the first protocells aggregating together into a progenote. Let us term the period of Earth’s history occupied by the self-assembly and evolution of progenotes the Progenean. As there is no rock record nor definable start or finish date, the Progenean cannot be used as a name for a geological era, but it must have been a distinct period in the emergence of biology. The mechanisms at play during life’s origins in the Progenean should impart a strong continuity with life’s long later evolution.
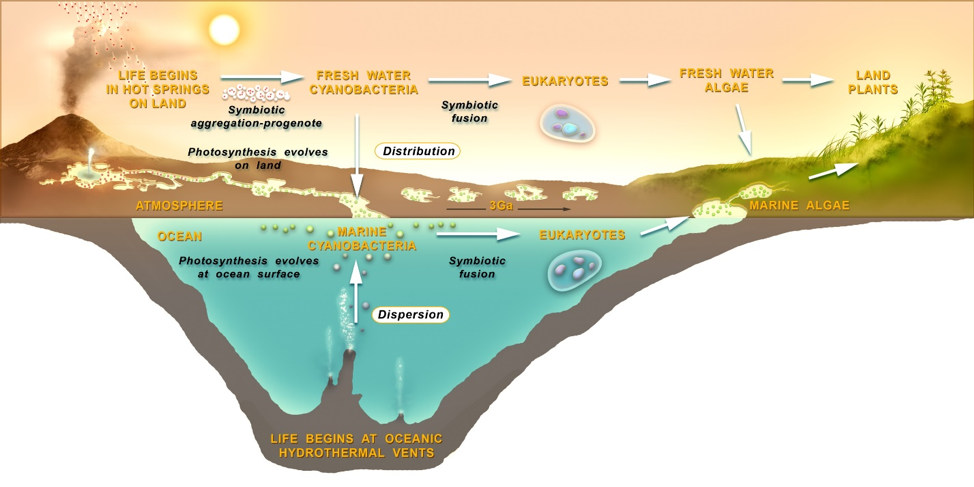
Figure 22. An origin of life and subsequent evolution in two alternative scenarios: at cycling hydrothermal pools on land or at hydrothermal vents deep in the ocean.
Placing two alternative hypotheses for an origin and subsequent evolution of life into one view (figure 22) we see that a Progenean origin of life on land (upper left) begins in a set of cycling, interconnected pools through a selection process targeting aggregates of symbiotic protocells. The pools allow combinations of the products of molecular evolution to be distributed across the landscape supporting a stepwise accumulation and sharing of key innovations. Diverse resources and stresses available in different aqueous settings permit gradients to be climbed to increasing robustness. Adaptations provide access to increasingly extreme conditions such as dilute streams and lakes, and eventually to osmotically challenging salty marine margins. Continuity is clearly expressed through the transformation of simple protocell aggregates into another more complex yet similar form: communities of living cells. Close aggregations of cells support pre-adaptations which prefigure the later rise of eukaryotes: larger cells which have through symbiotic fusion absorbed smaller ones. Figure 22 (center, right) depicts the long march of microbial life, now extant both in the oceans and in freshwater forms on land, to the rise of eukaryotic algae, fungi, and the first land plants and animals. Looking in reverse we can now appreciate the critical role of aggregation in the key transitions of evolution. These first protocell aggregations are a direct expression of niche construction so, we might conclude that niche construction underlies and is in fact a precondition for this first major transition from non-life to life.
Peering through the lens of an ocean origins, the situation looks far murkier. The oceans are a dispersing environment, unable to concentrate chemicals sufficiently to permit prebiotic chemistry. Therefore, the organic compounds provided by meteoritic in-fall to land-based pools are lost in the dilute bulk of the oceans. At deep sea hydrothermal vents, an alternative site proposed for life’s origins (Corliss et al. 1981; Russell et al. 2014), energy gradients and some chemical sources exist but in order to encapsulate (and concentrate) chemical reactions, pores in the minerals of the vent chimneys or gels composed of minerals have been suggested (Westall et al. 2018). However, if an interesting prebiotic chemical system was to emerge in one mineral pore or gel, it would remain entrapped there. Mineral compartments of any kind are not the same as membranous enclosures. Any chemical system must be able to engage in the vast cycles of combinatorial selection needed to drive evolution, and mineral pores or gels cannot be easily generated and subject to selection in these numbers. While no laboratory validation has been demonstrated with vent mineral encapsulation, lipid vesicles form naturally in pools on land in great numbers from meteoritic lipid sources and then easily encapsulate polymers. This has recently been empirically demonstrated in actual hot spring environments (Deamer et al. 2019). Lipid-bounded protocells are also impossible in the deep salty oceans as lipid vesicles collapse and crystallize in sea water (Milshteyn et al. 2018). If any form of deep ocean Progenean proto-life was dispersed far from the chemically rich vent environment it would immediately be starved in the dilute, cold bulk of the surrounding water. If this proto-life did manage to make it to the ocean’s surface where it could gain access to the other major available source of energy, sunlight, it would have to arrive with pre-adapted photosynthetic machinery in place. This is a highly implausible scenario. It may be argued however that life might originate in shallow hydrothermal vents where sunlight can still penetrate (up to 10m depth for instance) but the thermodynamic barriers to polymer and protocell formation within a degradative and dispersing salty sea water environment are still formidable. The dispersive environment of the oceans works against chemical concentration and cycling, and the essential steps of protocell formation, aggregation and combinatorial selection made possible by pools on land. The rise of eukaryotes is a process of symbiotic fusion which may also be strongly tied to pre-adaptations supported by aggregation and fusion supported by the land-based scenario. Of course, eukaryotes, fungi and the first animals could well have arisen in a marine shoreline environment, where nutrients from rock weathering still drive rich ecosystems today. The nutrient-poor open oceans have always been veritable deserts for life. Finally, ocean-originated life’s transition to land plants would have to go through the additional stage of the adaptation of marine algae to the freshwater environment (figure 22, upper right). As plants may have been shown to arise from a common ancestor living in a freshwater environment on land (Del-Bem, 2018), the old picture of “life crawling or growing out of the sea onto lifeless continents” seems outmoded.
In summary, a picture is recently emerging that life most plausibly got its start in hot little cycling pools possessing an ideal, “Goldilocks” prebiotic chemistry (Powner and Sutherland, 2011). Its subsequent adaptive evolution followed a natural downhill distribution across a landscape to more extreme environments of dilute rivers, lakes and then the salty sea shore. The open oceans and deep hydrothermal vents would have been colonized much later as they would have been even more extreme settings for early microbial communities.
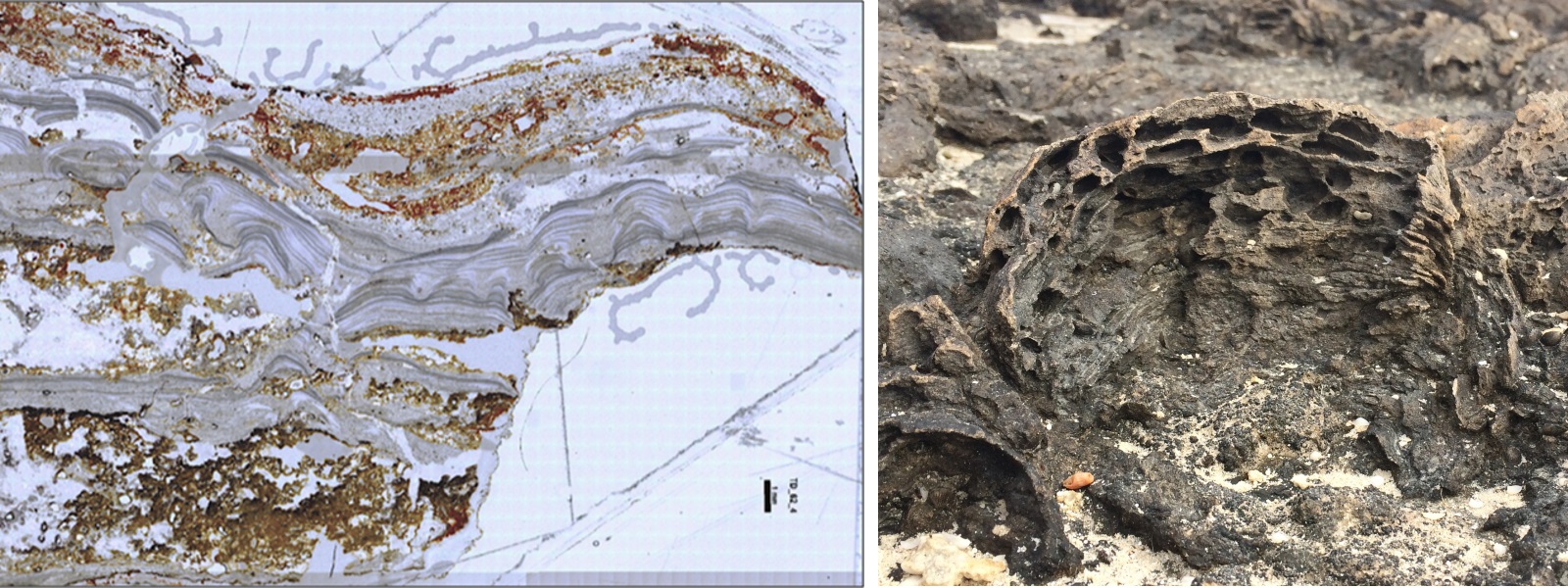
Figure 23. Left: geyserite preserving stromatolites indicative of microbial communities in a hot spring setting on land 3.5 billion years ago. Right: stromatolites newly formed at the marine shoreline of Shark Bay, Australia. Photo: Bruce Damer.
Lastly, it has been established that the oldest preserved evidence of microbial communities, laminar imprints in the rock called stromatolites (Djokic et al., 2017; Van Kranendonk et al., 2017), were present on land as far back as we can look, and whose living descendants today leave morphologically identical signatures (figure 23, left and right). With the accumulating experimental and geological evidence for an origin of life on land, and seemingly insurmountable thermodynamic barriers to the necessary prebiotic chemistry occurring in the oceans, it would seem that an origin of life needs to be taken seriously and resources expended to test the various parts of the hypothesis.
The hot spring hypothesis and the search for life in the universe
This “sea change” in the view of life’s origins carries major implications for our understanding of where it might arise elsewhere in the universe. The hot spring hypothesis described in these essays was used to support landing site selection for the Mars2020 rover (figure 24, left), a mission whose primary focus is the detection of evidence for past life on the surface of Mars. I was part of the team promoting a return to Columbia Hills, where an outcrop named Home Plate where rocks explored by the Spirit rover in 2007 turned up evidence of silica deposits, indicating that hot springs were operating in the area billions of years ago. Imaging of rocks revealed features (figure 24, right) resembling hot spring biosignatures at El Tatio in Chile (Ruff and Farmer, 2016). It emerged that an ancient hydrothermal spring resembling those on Earth had been discovered. Such environments are rife with life anywhere they are found on Earth and are excellent places to preserve evidence of that life for billions of years. While NASA eventually opted for another landing site location for Mars2020, the Columbia Hills site had advanced into the final selection round partly on these arguments. The team led by Arizona State University geologist Steve Ruff made the case that such hot spring deposits might be where life could get started in its “first outpost” on Mars. As the planet lost its atmosphere and surface water bodies due to the shutting down of its magnetic field, these springs might become a “last outpost” for life able to survive at the surface. As Mars lost its surface habitability microbial life could have escaped to wet and salty rock refuges protected from surface radiation. These microbes could have occasionally been brought back to the surface as volcanic hot spring environments erupted forming temporary oases. So, such ancient hot spring sites could be good places to look for evidence for life throughout Mars history.

Figure 24. Left: Mars2020 rover. Right: nodular silica structures imaged by the Spirit rover at Columbia Hills in 2007. Photo: NASA.
The search for life is also guiding NASA and other agencies to look to the icy moons of the outer solar system. The Cassini mission to Saturn detected, and flew through plumes spraying ice crystals through cracks in the ice at the south pole of the tiny moon Enceladus. It has been suggested that as life might start and thrives today at deep sea hydrothermal vents on Earth, then perhaps oceans under the ice shell of Enceladus might harbor hydrothermal activity, and living organisms. In their planning for the next mission to Enceladus and other icy moons, NASA scientists are using the approach that for life to start it needs land masses exposed to atmosphere as the “null hypothesis” for finding evidence for life there. In other words, without wet-dry cycling, Enceladus could be “habitable but sterile” and a mission to fly through the plumes would detect no clear evidence for life there.
Peering down into the short lens: P-I-M, three underlying properties of the living world
In the previous section we looked back through the long lens to life’s origins four billion years ago and peered out into the cosmos for clues as to where it might arise elsewhere. Next, we will peer down the short lens to the microscopic level. By diving deeply into the action of the progenote we may gain insights into the fundamental processes that can lift life from the background of purely physical universe.
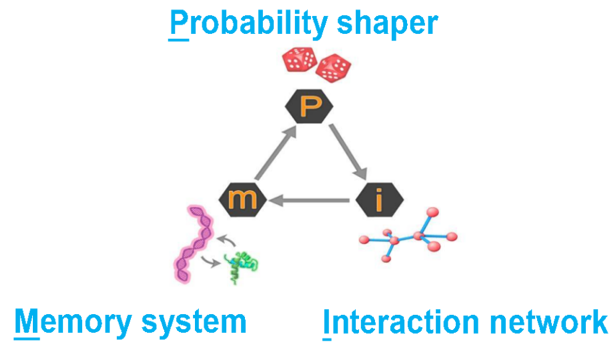
Fig 25. Abstracted version of the chemical cycle in the hot spring hypothesis scenario.
Figure 25 depicts an abstracted representation of the chemical cycle operating within a protocell and collectively within an aggregate of protocells (the progenote). A protocell is a means to crowd molecules close together so that reactions occur between them that would be highly unlikely dispersed in a dilute pool. Therefore, protocells are mechanisms to shape probability (P). When increasingly stable protocells aggregate into a progenote, a “network effect” can emerge in which interactions (I) occur through the diffusion of chemical products between protocells. Next, large numbers of polymers are cycled through combinatorial selection so that a few emerge possessing useful functions. Essentially, this process allows the cycling system to “discover” how to “write” polymer programs from initially random sequences. Some of these programs can act as a memory (M) permitting the replication of sets of polymers.
We propose that this abstract three-way Probability-Interaction-Memory (P-I-M) process, is a sort of engine of emergence, representing a universal mechanism for the origin and evolution of biology, and by extension human intelligence, culture and technology. This system could be generalized into a useful theoretical framework by which natural selection works on any system capable of supporting the three phases. The chemical scenario for the origin of life we have proposed here is only one such system.
Implications of the hot spring hypothesis for the Extended Evolutionary Synthesis (EES)
The hot spring hypothesis scenario proposes a set of mechanisms to transform matter from an inanimate to an animated state. It is also an experimentally-accessible exploratory system which could provide insights into the fundamental operating principles of evolution itself. For future thinking, we summarize the four main mechanisms the scenario employs:
- Membranous vesicles provide crowding to shape probabilities analogous to loading the dice, a network effect of interacting polymers in protocell aggregates which is the first self-constructed niche, Darwinian selection arises from this supporting niche, possibly through a proto-genetic memory template system arising through the formalism of P-I-M;
- Communities of dividing living cells emerge from earlier self-assembling transitory protocells;
- The first proto-genotypes arise through combinatorial selection of proto-phenotypes overcoming environmental hurdles;
- The vertical descent of the conserved genes of biological evolution originates from a system of horizontal sharing of the proto-genes resulting from chemical evolution.
If the protocell collective, the progenote, is the deepest ancestor of the microbial community, it must have arisen as a collaborative network working together to construct the mutually supportive environment necessary to cope with the stresses of the environment. When one progenote colony drifted into a pool occupied by another, a physical fusing of protocellular structures and sharing of innovations would have constructed the conditions in which a more robust “offspring” could emerge. While competition within and between microbial communities arose later, the initial steps of evolution would have been based entirely on more amorphous forms of recombination and sharing, mediated by proto-niche construction. This may give us some clues as to the balance between evolution marked by vertical descent of genes of organisms competing and models of symbiosis which also drive evolution today. In order for life to emerge, the initially passive proto-niche constructing processes of aggregating protocells had to evolve into active niche constructing processes of true living cells. The order of emergence of these active processes is experimentally testable and is speculated to include: structural polymers, pores, simple proto-genetic templates, metabolic cycles and catalysts, feedback controls, replicating mechanisms and, finally, protocell compartment division (Damer and Deamer, 2015; Deamer, 2019).
Seeking a specific link between life’s origins and its subsequent evolution we presented the hot spring hypothesis scenario to the Extended Evolutionary Synthesis community at the Evolution Evolving conference held at Churchill College, Cambridge on April 1-4th, 2019. Subsequent discussions with Richard Watson of Southampton University were most illuminating.
Drawing from new thinking by Watson and Thies in (Uller and Laland, 2019) they state that:
Niche construction and plasticity also have a more complex role in the individuality of the collective; the development of the collective phenotype. The need for multiple functional roles within a collective can be partially satisfied by simple compartmentalization (i.e., well-mixed within each collective) and stochastic differentiation (i.e., each particle adopts a phenotype probabilistically).
Considering our model of the emergence of a protocell-aggregate “progenote” as a proto-niche construction, in the language presented above it is made possible by the self-assembly through compartmentalization of a collective phenotype. That phenotype has a simple collective functional role for all “particles” which in our scenario are protocellular compartments with their encapsulated polymers. The first phenotypic role is to enhance the stability of the aggregate such that polymers are preserved and able to be elongated, re-synthesized and re-processed in a subsequent dry-wet cycle. Thus, the system starts out as un-differentiated and dominated by stochasticity but roles would then emerge as well as some internal organization as the progenote populations cycle through selection barriers. The process that drives this emergence begins with the initial accumulation of phenotypic traits selected from the random background supporting the collective into which then arises genotypic polymers to template future phenotypic polymers.
Watson and Thies go on to clarify that:
The co-construction of localized conditions (e.g., axial gradients, tissue differentiation, organogenesis, cell-cell signals etc.) that induce the right kind of plastic response (phenotypic form or function, abundance, timing, or movement, etc.) at the particle level is what we mean by development (of the collective phenotype).
Drilling into this second statement, perhaps the proto-niche construction we propose as the first scenario for development is expressed both through the physical construction of the niche (protocell aggregate), but also through signals (diffusion of products from one protocell in the aggregate to another causing a collective network effect). Perhaps our progenote can therefore express an early form of plasticity response as its collective phenotype and proto-genome adapts to selection pressures. Initially entirely phenotypic the in the early phases, the progenote adopts a diffuse genotype and over time the particles represented by sets of interacting phenotype-genotype polymers become coherently associated within protocell compartments. Those compartments grow their role-identity until the physical budding mechanism of protocell propagation is replaced by controlled division leading to the transition to “living” cells. None of this development is possible for separate, free-floating protocells, and can only occur in the environment of the proto-niche of the progenote protocellular mass. So, it may be plausible to state that “niches came first.” However, to clarify, the proto-niches are also themselves impermanent as the separation of the aggregate into temporary individual protocells budded off during rehydration of lamellae is an essential step to subject protocells to individual selection and drive their robustness. Thus, the scaffold of a cycling chemical system in a hot little pool provides distinct phase changes with the distinct physical processes of each phase replaced by molecular machinery in later living cells.
The linking together of this newly proposed hypothesis of life’s origins with the its subsequent evolution is only just beginning. We hope that a continuing exercise studying and experimentally testing this scenario, situated at the simplest starting point of life, will provide insight into its later highly complex and sometimes opaque evolutionary processes.
Considerations for the first major transition in evolution
We would also like to propose that the construction of the proto-niche of the progenote, and the scaffold it provides for the stepwise evolution of the functions of dividing, living cells, is the first major transition in evolution. Maynard Smith and Szathmáry (1995) identified several properties common to these transitions. We can relate the progenote and the above origin scenario to each of these properties:
- Smaller entities have often come about together to form larger entities. Protocells aggregate into a larger entity, the progenote.
- Smaller entities often become differentiated as part of a larger entity. Polymer populations complexify as evolutionary adaptations appear in a stepwise fashion partly driven by their participation in the network operating in a progenote.
- The smaller entities are often unable to replicate in the absence of the larger entity. Protocells cannot divide on their own when suspended in solution but can replicate through the synthesis and elongation of sets of polymers within the progenote, which then deposits them into new protocells which bud off during rehydration.
- The smaller entities can sometimes disrupt the development of the larger entity. Evolution of new functional polymers with catalytic abilities may disrupt and replace the functions of other protocells or entire operating networks within a progenote aggregate.
- New ways of transmitting information have arisen. Over many cycles, initially freely mixed short proto-genetic sequences become associated into longer strands which express more functions and become associated with individual protocells until they support the machinery of more robust living cells capable of division. This new form of genetic information is conserved by cells and passed down to descendants.
Summary
We have now taken a trip back four billion years to a 21st Century science version of Charles Darwin’s warm little origin pond. We found ourselves at the edge of a hot little cycling pool connected to a hydrothermal spring on a volcanic landscape. Using the boot-strap metaphor from computer science we found that our pool cycles between wet and dry states and can boot-strap a molecular operating unit called a protocell capable of subjecting sets of polymers to chemical evolution. We then proposed the transitional entity that carried inanimate matter across the vast evolutionary chasm to the living world: the progenote. The properties of that entity, first arising through a natural process of self-assembly we term proto-niche construction is powered by another computing metaphor, the network effect. Next, we considered how through the principle of continuity the progenote initiates and supports the first and subsequent major transitions in evolution, through aggregation, to fusion supporting the endosymbiotic merger of eukaryotes, and the emergence of communal multi-cellular life such as fungi, algae, plants and animals. As a new branch of science gets underway which is dedicated to re-creation of the progenote in the laboratory, we predict that there will be many important new discoveries and implications for evolutionary theory and humanity.
So, which came first, the chicken or the egg?

Figure 26. Which came first, the chicken or the egg?
Arriving at the end of our journey have we become enlightened enough to address the age-old question: which came first, the chicken or the egg? (figure 26). Perhaps the protocells and their aggregation into the progenote, able to arise through nothing more than self-assembly processes within the kinetic trap of wet-dry cycling pools, represents the first “chicken.” The progenote can then undergo a form of growth by “feeding” on extra monomers, lipid and other available compounds in the concentrating phase of the drying pool. In its early evolutionary cycles, the progenote can express functions that are replaced by later active processes in biology. So, in this way we could say that the progenote expresses a kind of phenotype or emergent chicken. But the progenote does not yet possess the ability to create an “egg”, a simple proto-genetic template to help generate future progenotes, specifically replicating individual protocells with their sets of cycling polymers. As we suggested in the above abstracted P-I-M model, the shaping of probability to enable rare events to occur, followed by the network effect set up within an aggregate of protocells is postulated to form a substrate upon which some form of templated replication might arise. This is solidly in the realm of speculation but early experiments by our group and others suggest that this is plausible. So, in the end, a primitive proto-chicken arises, learns to scratch a meagre existence in its chicken yard, shares its amorphous boneless parts with other chickens over a long period of time until one day a group of chickens is able to lay the first egg. In the end, or perhaps in the beginning, it is neither the chicken nor the egg that came first, but it is instead the collection of chicken yards, possessing just the right combinatorial conditions for both chickens and eggs to be cycled into existence.
One final thought: should we re-factor the platform of evolution?
Out beyond today’s complex web of life of innumerable chickens and eggs, lies something greater, a meta-process. By unwinding life to its starting point, we can more clearly identify this process. The First protocells are entirely dependent on the wet-dry cycling pools and their external inputs of energy and building blocks. Systems of interacting polymers are selected from random sequences and can evolve only within the supporting matrix of an aggregate of protocells called the progenote. Through many selective cycles, distribution and cross-sharing between progenote aggregates and a prototypical form of niche construction begins. A simple proto-genetic function – templated replication – can then emerge, supported by these niches. A gene-centric view of life would have the replication of polymers emerge first and drive the process of evolution. Yet genes today cannot exist or replicate without the support system of their surrounding cell. The de novo emergence of a system of replicating polymers without such a support system is chemically implausible. Therefore, niches must have come first.
The hot spring hypothesis scenario and empirical evidence behind it presented in this four part essay suggests how evolution originated and predicts the nature of the first evolving chemical systems and their path to living cellular communities. This path begins with a primordial form of niche construction, adds a network effect, adaptation through distribution across a landscape, and a resulting stepwise evolution of polymer systems which ultimately combine to support the transition into the first living cellular communities. The meta-system which enables this process is the aggregation of simpler individuals, sharing of resources across aggregations, and growth and evolution at the level of the aggregation to drive increasingly robust innovations within the individuals. After rise of living cellular forms within these aggregations they remain dependent and indivisible from their surrounding microbial communities. Evolutionary theory has historically rested heavily on the study of animals, interpreting their behavior as individuals competing “red in tooth and claw” to pass on their genes. Yet from its origin and for most of its evolutionary history, life has been dominated by niche construction and networks of resource sharing between collaborating organisms, primarily microbial consortia. Animals are completely dependent on grazing on the products of these consortia to act out the drama of their lives. It therefore behooves science to consider that perhaps life, and evolution itself, actually rests on a platform more broadly based on niches and collaborative networks than the strict competition between individuals struggling to pass on their genetic lines.
References for the series
• Arnoldt H., Strogatz S., Timme, M. (2015) Toward the Darwinian transition: Switching between distributed and speciated states in a simple model of early life. Phys. Review E 92, 052909.
• Bartel D. and Szostak J. (1993) Isolation of new ribozymes from a large pool of random sequences. Science, 261, 1411–1418.
• Corliss J., Baross J., Hoffman S. (1981) An hypothesis concerning the relationship between submarine hot springs and the origin of life on Earth. Oceanol Acta 4: 59-69.
• Damer B. (2016) A field trip to the Archaean in search of Darwin’s warm little pond. Life.
• Damer B. and Deamer D. (2015) Coupled phases and combinatorial selection in fluctuating hydrothermal pools: a scenario to guide experimental approaches to the origin of cellular life. Life (Basel) 5: 872-887.
• Damer B., and Deamer D. (2019) The hot spring hypothesis for an origin of life, Astrobiology, in preparation.
• Darwin C. (1859). On the origin of species by means of natural selection, or preservation of favoured races in the struggle for life. London: John Murray,
• Darwin C. (1871) Darwin Correspondence Project, “Letter No. 7471”, Available online: https://www.darwinproject.ac.uk/letter/?docId=letters/DCP-LETT-7471.xml (accessed on 20 March 2019).
• Deamer D. (2019) Assembling Life, How Can Life Begin on Earth and Other Habitable Planets? Oxford University Press.
• Deamer D. and Pashley R. (1989) Amphiphilic components of the Murchison carbonaceous chondrite: Surface properties and membrane formation. Orig. Life Evol. Biosphere 19:21-38.
• Deamer D., Damer B., Kompanichenko V. (2019) Hydrothermal chemistry and the origin of cellular life. Astrobiology in press.
• Del-Bem L.E. (2018) Xyloglucan evolution and the terrestrialization of green plants. New Phytologist. 219 (4): 1150–1153
• Djokic T., Van Kranendonk M., Campbell K., Walter M., Ward C. (2017) Earliest Signs of Life on Land Preserved in ca. 3.5 Ga Hot Spring Deposits. Nat Commun 8: 16149.
• Dyson F. (1999) Origins of Life, Cambridge University Press.
• Hargrave M., Spencer S., Deamer D. (2018) Computational Models of Polymer Synthesis Driven by Dehydration/Rehydration Cycles: Repurination in Simulated Hydrothermal Fields. J Mol Evol.
• Higgs P. (2016) The effect of limited diffusion and wet-dry cycling on reversible polymerization reactions: Implications for pre biotic synthesis of nucleic acids. Life (Basel)..
• Maynard Smith J. and Szathmáry E. (1995). The Major Transitions in Evolution. Oxford, England: Oxford University Press. ISBN 978-0-19-850294-4.
• Miller S.L. (1953) A Production of Amino Acids under Possible Primitive Earth Conditions, Science, Vol. 117, No. 3046, pp. 528-529.
• Milshteyn D., Damer B., Havig J., Deamer D. (2018) Amphiphilic compounds assemble into membranous vesicles in hydrothermal hot spring water but not in seawater. Life (Basel).
• Odling-Smee J., Laland K., Feldman M. (2003) Niche Construction: The Neglected Process in Evolution. In Monographs in Population Biology; Princeton University Press: Princeton.
• Orgel L. (1968) Evolution of the genetic apparatus. Journal of Mol. Biology, Vol. 38, Iss. 3, 28, pp. 381-393
• Pearce B., Pudritz R., Semenov D., Henning T. (2017) Origin of the RNA World: The fate of nucleobases in warm little ponds. Proc Natl Acad Sci USA
• Powner M. and Sutherland J. (2011). Prebiotic chemistry: a new modus operandi. Philosophical transactions of the Royal Society of London. Series B, Biological sciences, 366(1580), 2870-7.
• Rajamani S., Vlassov A., Benner S., Coombs A., Olasagasti F., Deamer D. (2008). Lipid-assisted synthesis of RNA-like polymers from mononucleotides. Orig Life Evol Biosph 38:57-74.
• Ross D. and Deamer D. (2016) Dry/Wet cycling and the thermodynamics and kinetics of prebiotic polymer synthesis. Life (Basel) doi: 10.3390/life6030028.
• Ruff S.W. and Farmer J.D. (2016) Silica deposits on Mars with features resembling hot spring biosignatures at El Tatio in Chile. Nat. Commun. 7, 13554.
• Russell M., Barge L., Bhartia R., Bocanegra D., Bracher P., Branscomb E., Kidd R., McGlynn S., Meier D., Nitschke W., Shibuya T., Vance S., White L., Kanik I. (2014) The drive to life on wet and icy worlds. Astrobiology 14: 308 -343.
• Sawai H., Lohrmann, R., Orgel, L (1975) Prebiotic peptide-formation in the solid state. J. Mol. Evol. 6: 165.
• Uller T. and Laland K.N. (2019) Evolutionary Causation. Biological and Philosophical Reflections. MIT Press: Cambridge, Mass. In Press.
• Van Kranendonk M., Deamer D., Djokic T. (2017) Life Springs: Life on Earth Came from a Hot Volcanic Pool, Not the Sea, New Evidence Suggests. Scientific American 317(2):28-35.
• Westall F., Hickman-Lewis K., Hinman N., Gautret P., Campbell K.A., Bréhéret J.G., Foucher F., Hubert A., Sorieul S., Dass A.V., Kee T.P., Georgelin T., Brack A. (2018) A Hydrothermal-Sedimentary Context for the Origin of Life. Astrobiology 18:3, 259-293.
• Woese C. (1998) The universal ancestor. Proc Natl Acad Sci USA, 12: 6854-6859.
• Woese C. (2002). On the evolution of cells. Proc Natl Acad Sci USA, 99: 8742–8747.
• Woese C. and Fox G. (1977) The concept of cellular evolution. J Mol Evol 10: 1-6.
Additional Media Resources
In the beginning: The origin and purpose of life
A 2015 TEDx talk by the author.
Local Focus: American scientist researches the origins of life in Rotorua
Video by New Zealand Herald about the author’s June 2018 field work in the hot springs at Rotorua.
Living Universe: Origin of life fieldwork at Bumpass Hell
Living Universe documentary segment about field work by David Deamer and the author in the fumarole vents of Bumpass Hell, California in August 2016.
The origin of life and the future of consciousness
Broader societal and philosophical implications and speculations presented by the author at Science and Nonduality 2017.