Does Inheritance Need a Rethink? Conceptual Tools to Extend Inheritance beyond the DNA
by Sophie Veigl, Javier Suárez, & Adrian Stencel
5 November 2019
Inheritance is an essential component of evolutionary processes. Without inheritance, evolution by natural selection cannot lead to the accumulation of complex, adaptive traits. But what can be inherited, and how?
Eva Jablonka and Marion J. Lamb argued in “Evolution in Four Dimensions” that in addition to genetic inheritance, evolution can also work on epigenetic, behavioral, and symbolic/linguistic forms of inheritance. While this proposal seems appealing, it is generally assumed that only genetic inheritance, especially “DNA-based” inheritance, is faithfully replicated within lineages. From this traditional standpoint, traits, behaviors, and extra-genetic materials such as cellular content, microbes, or environments, are only heritable because they have a genetic basis. This stance lends genetic inheritance a privileged status. However, new data revealing the scale and diversity of extra-genetic forms of inheritance have increasingly led to the traditional stance being questioned.
This year at the biennial meeting of the International Society of the History, Philosophy, and Social Studies of Biology (ISHPSSB) in Oslo, we– Sophie Veigl, Javier Suárez, and Adrian Stencel — got together to consider in what ways a “DNA-based” conception of biological inheritance can hamper the proposal of alternative accounts of biological inheritance. We discovered common threads in each of our respective areas of expertise (small RNA inheritance, holobionts, niche construction) that point to alternative conditions for inheritance.
We found that a central “pressure” to adhere to DNA-based accounts comes from two required conditions for inheritable entities:
1) the replication of entities that carry information, and
2) the stability of these entities throughout generations.
While it is, of course, possible that alternative forms of inheritance involve entities that are replicated and transgenerationally stable, these conditions are not necessary for evolutionarily-relevant inheritance. We should not expect that alternative trajectories of inheritance fulfil these requirements. Those advocating for a more inclusive approach do not need to argue that their forms of inheritance are as stable or faithful as genetic inheritance.
Alternatively, we define the Stability of Traits (SoTs) as the phenomenon by which a phenotypic trait persists throughout generations in a population. Reconstitution are the processes through which SoTs is transgenerationally realized in a population.
In DNA-based inheritance, SoTs is satisfied because a specific entity (i.e., DNA) causally related to the phenotype is replicated and transmitted from the parents to the offspring. In the offspring, the entity is causally related to the phenotype.
In our case studies, SoTs is realized through different processes of reconstitution that do not require the direct transmission of the entity that causes the phenotype. The concept of “reconstitution” goes beyond the much narrower definitions of stability and faithful replication that is only satisfied by DNA-based inheritance. Reconstitution applies to a vast range of processes, across different “levels” or “scales.”
In the following, we examine reconstitution as a concept describing a) the inheritance of a specific molecular entity, b) the holobiont as a unit of selection and c) niche construction.
Small RNAs (Sophie Juliane Veigl)
In our first case study, we approach a specific type of epigenetic inheritance: the inheritance of small RNAs. Small RNAs can store information, are amplifiable, and responsive to environmental stimuli. Small RNAs are short molecules that target and destroy specific messenger RNAs (mRNAs), thereby inhibiting the synthesis of a specific protein.
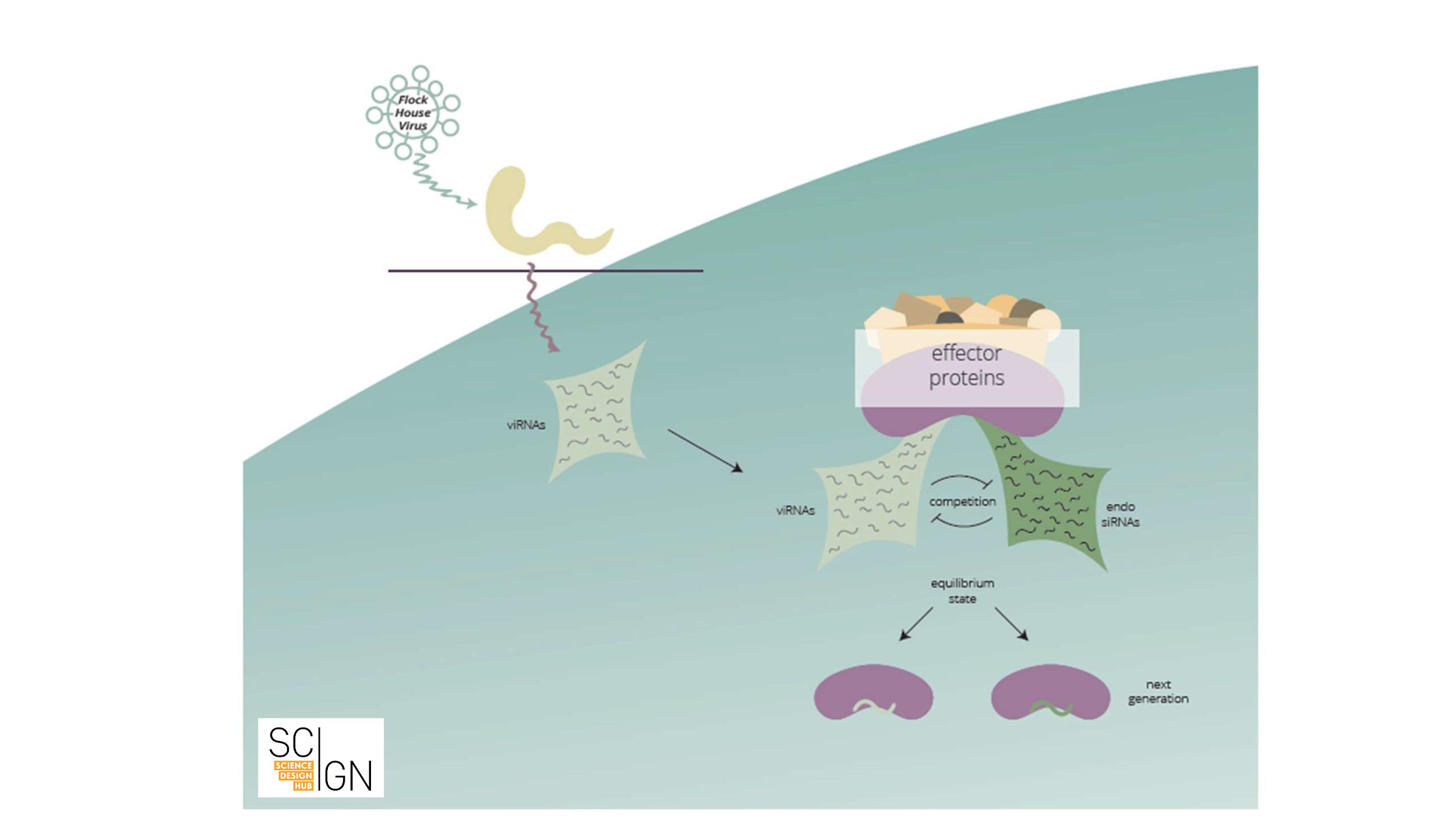
Let us provide a brief example: Small RNAs were shown to target and destroy viral RNAs. Small RNAs storing the information of which viral RNAs to destroy are amplified and inherited. In a sense, future generations are thus “immunized” against a specific virus. Thus, an adaptive small RNA response against an environmental stimulus can be transgenerationally maintained (see Fig. 1).
Figure 1 Viral infection elicits a small RNA-based response in C. elegans (scheme). Anti-viral RNAs (viRNAs) that are complementary to viral RNA are synthesized upon viral infection. These viRNAs compete with endogenous small RNAs for binding to small RNA effector proteins, that mediate transgenerational inheritance of small RNAs. As a result, viRNAs are inherited and confer viral immunity to subsequent generations.
Critics of small RNA inheritance usually raise two pressing conceptual problems.
1) How can the effects of small RNAs be maintained throughout generations?
2) How can a specific small RNA be faithfully transmitted?
These questions, however, commit the fallacy of presupposing a DNA-based perspective: they insist on the replication of entities and their stability throughout generations. But should we expect alternative trajectories of inheritance to fit a “DNA-based” conceptual toolbox?
The first conceptual problem is linked to the maintenance of chromatin modifications. Small RNAs are responsible for the deposition of chromatin modifications. Vice versa, the amplification of small RNAs (in each generation) is dependent on specific chromatin modifications. But chromatin modifications were shown to get erased during generational checkpoints. It is thus not clear how chromatin modifications or small RNAs can be maintained transgenerationally. Is it still possible to ensure replication?
While probably possible, replication is not necessary. Imagine a scenario in which small RNAs reconstitute chromatin modifications in each generation. In so doing, they reconstruct the gene regulation of a past generation. At the same time, by reconstituting the chromatin landscape of the past, small RNAs promote their own amplification. In this scenario, chromatin modifications need not be replicated: they are reconstituted. Phenotypic effects will be the same. In conclusion, replication of a specific causally relevant entity is not required to maintain a trait transgenerationally.
Shifting the focus to small RNA-based reconstitution invokes a second problem: small RNA maintenance. Tens of thousands of small RNAs are contained within each cell. If one specific type of small RNA (e.g., that storing the information of a viral RNA sequence) is causally relevant for a phenotype, how can it travel from one generation to the next? Ensuring the stability of one specific small RNA entity poses an insurmountable administration and delivery problem. There is probably no way to ensure faithful delivery/distribution and maintenance of a specific small RNA. Thus, the stability of entities cannot be ensured. But it seems not to be required either.
Although the stability of a specific small RNA entity cannot be ensured, we are nevertheless able to observe the stability of relevant traits caused by small RNAs. This invokes the necessity of a major rethink in small RNA inheritance. Instead of asking: “How can a specific molecule be maintained?” we need to ask: “Which conditions need to be met in order to guarantee the stability of traits?”
Holobionts (Javier Suárez)
In our first case study we argued that small RNA inheritance cannot be explained through the concepts of “stability of entities” and “replication”. We will now show the same for two more case-studies, the first being the case of holobionts.
Holobionts are ecological entities composed of a macrobial host (currently confined to animal or plant) plus the set of associated microorganisms that compose its microbiome. Holobionts have recently been proposed to be units of selection and their hologenomes (host genome plus particular microbial genes that make up the microbiome) have being argued to reproduce as a sole unit (which are technically called “reproducers”). Holobionts are ubiquitous, as most animals and plants host microbes that have significant phenotypic effects on their fitness.
One problem of considering hologenomes as reproducers is that the species that constitute the microbiome of each host are not transgenerationally transmitted. In each generation, they are recruited (sometimes randomly) from the environment. Thus, it was argued that this disruption of the inheritance relations precludes that hologenomes are reproducers, and therefore that holobionts are not units of selection. In what follows we will challenge this perspective.
The criticism is based on a replication model of inheritance. Critics assume that hologenomes can only be reproducers if an essential subset of the species that compose the host microbiome is vertically transmitted to the next generation. They require that the inheritance of hologenomes operates in the same way in which DNA is vertically transmitted through replication. This model is not satisfied in most holobionts. And it doesn’t need to be satisfied.
Rather than thinking of hologenomes by appealing to the “DNA-based” conceptual toolbox, one must ask the following questions:
1) Can hosts effectively reconstruct their phenotypic traits transgenerationally?
2) If so, does the microbiome play a substantial role in this process?
3) What type of mechanisms might have evolved to guarantee microbiome-mediated trait preservation?
Contrary to the replication paradigm, the reconstitution paradigm requires that if the answers to 1) and 2) are affirmative for a given animal-microbiome system, despite the lack of microbiome replication, then we need an alternative explanation that accounts for the preservation of these traits (Fig. 2). Expecting that these explanations will satisfy the replication paradigm, or discarding any explanation that does not satisfy that paradigm, is not an acceptable scientific attitude.
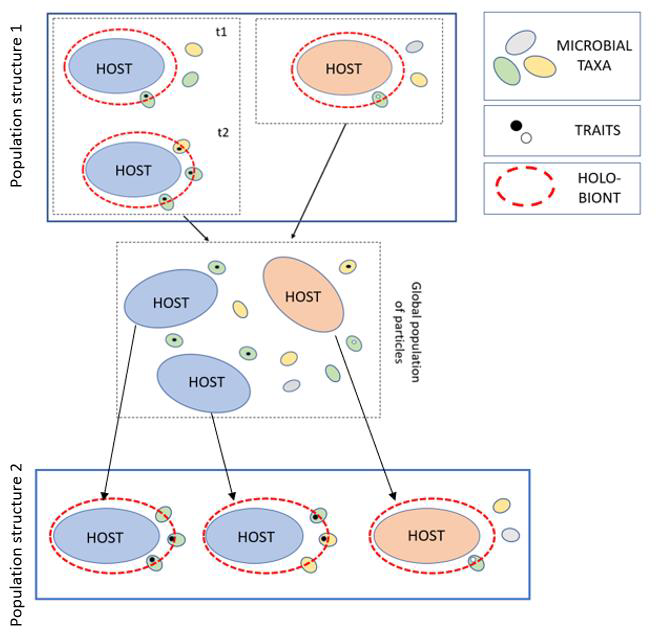
Figure 2. Application of the ideas of reconstitution and stability of traits to holobionts. The figure shows the evolution of a holobiont population through time, and how its structure changes transgenerationally. Different colours represent different genomes, both in the host and in the bacterial taxa.The coloured circles represent the bacterial contribution to different phenotypic traits. Finally, the arrows represent reproductive events. Reconstitution occurs in the global population of particles. Reconstitution among holobionts would theoretically give rise to a new population structure in which the blue host genome tends to be selectively assorted with bacterial taxa that bear the components that are required to develop the phenotypic trait (and thus, to cause the phenomenon of stability of traits), whereas the same is not the case with the red host genome. The figure shows a theoretical possibility in which we may have inheritance that does not satisfy the replication paradigm, and which we believe is the general case among holobionts. From Suárez 2019: 226, Figure 29.
Ecological inheritance (Adrian Stencel)
After having presented two case-studies that necessitate a rethink of the concepts employed for describing inheritance, we will bring home our point with one last case study on ecological inheritance. Ecological inheritance refers to the situation of inheriting resources and environmental conditions that were present during the parents’ lifetime. Resources that are ecologically inherited are often pivotal for keeping similarity in the parent-offspring line. They might be the causal factor responsible for the similarity observed. Let us consider a simple example to explain the importance of ecological inheritance.
Suppose that you have a particular heart disease and where you live, there is no available medical treatment. People with this condition cannot get sexual partners and reproduce because they die at the age of 12-14 years. One day you move to a place where the available medicine can help you to cope with your heart problems. You get cured, you find a partner, and reproduce. Your children have the same disease. As they live in a place where the disease can be cured, they can be treated, and their life can continue normally.
This story shows two things about ecological inheritance. First, it represents the most radical example of reconstitution, as the phenotypic trait – in our story, a healthy heart – is kept stable between generations. Stability only occurs because the offspring has access to a subset of resources (advanced medicine) that was inherited (as your children only had it because you had it), but not transferred from the parents. A particular phenotype depends on a complex subset of social interactions that can be used to re-constitute the phenotype of the parents.
Second, it shows that ecological inheritance is vulnerable to changes induced by the individual. Organisms can actively change the availability of resources and the selective pressures from the environment, having substantial impact on their ecological inheritance. This process is called niche construction. In the scenario above, the parents were capable of changing the availability of resources and the selective pressures by migration. Furthermore, the change in the selective pressure was inherited by their offspring.
One might argue that ecological inheritance is not proper heredity. But it is heredity, because it causes parent-offspring similarity in certain circumstances. Of course, critical questions are: to what extent is ecological inheritance responsible for phenotypic similarities? What is the difference between ecological inheritance and other channels of inheritance? These questions only make sense if we switch from the DNA-replication paradigm towards reconstitution. Instead of emphasizing a specific inheritance mechanism reconstitution asks which different mechanisms that ensure the stability of traits exist across different “scales” and “levels” of inheritance.
What now? Some prospects of the shift towards reconstitution for contemporary biological practice
Above we provided three examples in which “reconstitution” and “SoTs” describe phenomena much more adequately than do the concepts “replication” and “stability of entities”. It is important to note that the latter concepts are just particular ways of realizing SoTs and reconstitution. They are adequate for some instances of DNA-based inheritance but their realization is in no way a prerequisite for other inheritance processes. We believe that reconstitution and SoTs are a multiply realizable processes and multiply realizable phenomena. Different cases of SoTs involve processes of reconstitution other than replication.
Several clarifications are of course still required. To provide an outlook on the consequences of what was discussed above, we point out three advantages that the use of the concepts of SoTs and reconstitution bring to biological research:
1. “Reconstitution” and “SoTs” introduce an extended perspective on possible sources of phenotypic variation. Rather than assuming that DNA is the only source of phenotypic variation, SoTs and reconstitution encourage studying alternative sources of heritable variation.
2. The concept of reconstitution encourages us to discover new processes that guarantee SoTs. Such new process may be distinct in several ways: the organism/environment dichotomy will be less sharp than is usually assumed. Additionally, realization of these processes may be very diverse: they may be realized through entities, network-structures, concentration rates, etc.
3. Finally, “reconstitution” and “SoTs” make possible a more comprehensive view of the evolutionary process. The concepts of reconstitution and SoTs call for a different application of vital evolutionary concepts such as “natural selection,” “drift,” or “adaptation,” insofar as they call for an “enlargement” of the sources that can go through any of these processes.
Further Reading:
Bounduriansky, R., & Day, T. (2018) Extended Heredity, A New Understanding of Inheritance and Evolution. Princeton University Press.
Laland, K., Matthews, B. and Feldman, M.. (2016). An introduction to niche construction theory. Evolutionary Ecology 30: 191–202.
Laland, K., Uller, T., Feldman, M., Sterelny, K., Müller, G. B., Moczek, A., … & Futuyma, D. J. (2014). Does evolutionary theory need a rethink?. Nature News, 514(7521), 161.
Miska, E. A., & Ferguson-Smith, A. C. (2016). Transgenerational inheritance: models and mechanisms of non–DNA sequence–based inheritance. Science, 354(6308), 59-63.
Odling-Smee, J. and Laland, K. (2011). Ecological Inheritance and Cultural Inheritance: What Are They and How Do They Differ? Biological Theory 6: 220–230.
Rechavi, O., Minevich, G., & Hobert, O. (2011). Transgenerational inheritance of an acquired small RNA-based antiviral response in C. elegans. Cell, 147(6), 1248-1256.
Rechavi, O., & Lev, I. (2017). Principles of transgenerational small RNA inheritance in Caenorhabditis elegans. Current Biology, 27(14), R720-R730.
Sarkies, P., Ashe, A., Le Pen, J., McKie, M. A., & Miska, E. A. (2013). Competition between virus-derived and endogenous small RNAs regulates gene expression in Caenorhabditis elegans. Genome research, 23(8), 1258-1270.
Stencel, A. & Wloch-Salamon, D. M. (2018). “Some theoretical insights into the hologenome theory of evolution and the role of microbes in speciation”, Theory in Biosciences, 137 (2): 197-206.
Stencel A, Crespi B (2013) What is a genome? Mol Ecol 22:3437–3443.
Stencel A (2016) The relativity of darwinian populations and the ecology of endosymbiosis. Biol Philos 31:619–637.
Suárez, J. (2018) The importance of symbiosis in philosophy of biology: An analysis of the current debate on biological individuality and its historical roots Symbiosis 76/2: 77-96.
Suárez, J. (2019) The hologenome concept of evolution: a philosophical and biological study. PhD dissertation. University of Exeter. http://hdl.handle.net/10871/39349. Accessed 29/10/2019.
Suárez, J. & Triviño, V. (2019). A metaphysical approach to holobiont individuality: Holobionts as emergent individuals Quaderns de Filosofía 6/1: 59-76.
Veigl, S. J. (2017). Use/disuse paradigms are ubiquitous concepts in characterizing the process of inheritance. RNA biology, 14(12), 1700-1704.
Veigl, S. J. (2019). Seeing “Lamarckian” More Positively: The Use/Disuse Paradigm Increases Understanding. BioEssays, 1900054.